+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8598 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of human bocavirus 1 | |||||||||
 Map data Map data | Human bocavirus 1 (VP2 expressed in Sf9 cells using recombinant baculoviruses) | |||||||||
 Sample Sample | human bocavirus 1 != Human bocavirus 1 human bocavirus 1
| |||||||||
 Keywords Keywords | HBoV1 / human parvovirus 1 / parvovirus / respiratory tract infection / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationA2-type glycerophospholipase activity / phospholipase A2 / T=1 icosahedral viral capsid / lipid catabolic process / viral capsid / host cell cytoplasm / host cell nucleus / structural molecule activity Similarity search - Function | |||||||||
| Biological species |  Human bocavirus 1 Human bocavirus 1 | |||||||||
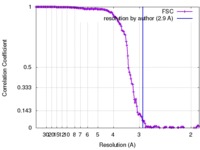
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Mietzsch M / Kailasan S | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2017 Journal: J Virol / Year: 2017Title: Structural Insights into Human Bocaparvoviruses. Authors: Mario Mietzsch / Shweta Kailasan / Jamie Garrison / Maria Ilyas / Paul Chipman / Kalle Kantola / Mandy E Janssen / John Spear / Duncan Sousa / Robert McKenna / Kevin Brown / Maria Söderlund- ...Authors: Mario Mietzsch / Shweta Kailasan / Jamie Garrison / Maria Ilyas / Paul Chipman / Kalle Kantola / Mandy E Janssen / John Spear / Duncan Sousa / Robert McKenna / Kevin Brown / Maria Söderlund-Venermo / Timothy Baker / Mavis Agbandje-McKenna /    Abstract: Bocaparvoviruses are emerging pathogens of the family. Human bocavirus 1 (HBoV1) causes severe respiratory infections and HBoV2 to HBoV4 cause gastrointestinal infections in young children. Recent ...Bocaparvoviruses are emerging pathogens of the family. Human bocavirus 1 (HBoV1) causes severe respiratory infections and HBoV2 to HBoV4 cause gastrointestinal infections in young children. Recent reports of life-threatening cases, lack of direct treatment or vaccination, and a limited understanding of their disease mechanisms highlight the need to study these pathogens on a molecular and structural level for the development of therapeutics. Toward this end, the capsid structures of HBoV1, HBoV3, and HBoV4 were determined to a resolution of 2.8 to 3.0 Å by cryo-electron microscopy and three-dimensional image reconstruction. The bocaparvovirus capsids, which display different tissue tropisms, have features in common with other parvoviruses, such as depressions at the icosahedral 2-fold symmetry axis and surrounding the 5-fold symmetry axis, protrusions surrounding the 3-fold symmetry axis, and a channel at the 5-fold symmetry axis. However, unlike other parvoviruses, densities extending the 5-fold channel into the capsid interior are conserved among the bocaparvoviruses and are suggestive of a genus-specific function. Additionally, their major viral protein 3 contains loops with variable regions at their apexes conferring capsid surface topologies different from those of other parvoviruses. Structural comparisons at the strain (HBoV) and genus (bovine parvovirus and HBoV) levels identified differences in surface loops that are functionally important in host/tissue tropism, pathogenicity, and antigenicity in other parvoviruses and likely play similar roles in these viruses. This study thus provides a structural framework to characterize determinants of host/tissue tropism, pathogenicity, and antigenicity for the development of antiviral strategies to control human bocavirus infections. Human bocaviruses are one of only a few members of the family pathogenic to humans, especially young children and immunocompromised adults. There are currently no treatments or vaccines for these viruses or the related enteric bocaviruses. This study obtained the first high-resolution structures of three human bocaparvoviruses determined by cryo-reconstruction. HBoV1 infects the respiratory tract, and HBoV3 and HBoV4 infect the gastrointestinal tract, tissues that are likely targeted by the capsid. Comparison of these viruses provides information on conserved bocaparvovirus-specific features and variable regions resulting in unique surface topologies that can serve as guides to characterize HBoV determinants of tissue tropism and antigenicity in future experiments. Based on the comparison to other existing parvovirus capsid structures, this study suggests capsid regions that likely control successful infection, including determinants of receptor attachment, host cell trafficking, and antigenic reactivity. Overall, these observations could impact efforts to design antiviral strategies and vaccines for HBoVs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8598.map.gz emd_8598.map.gz | 81.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8598-v30.xml emd-8598-v30.xml emd-8598.xml emd-8598.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8598_fsc.xml emd_8598_fsc.xml | 16.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_8598.png emd_8598.png | 297 KB | ||
| Filedesc metadata |  emd-8598.cif.gz emd-8598.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8598 http://ftp.pdbj.org/pub/emdb/structures/EMD-8598 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8598 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8598 | HTTPS FTP |
-Related structure data
| Related structure data |  5urfMC  8604C  8605C  5us7C  5us9C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8598.map.gz / Format: CCP4 / Size: 242.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8598.map.gz / Format: CCP4 / Size: 242.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human bocavirus 1 (VP2 expressed in Sf9 cells using recombinant baculoviruses) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.95 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human bocavirus 1
| Entire | Name:  human bocavirus 1 human bocavirus 1 |
|---|---|
| Components |
|
-Supramolecule #1: Human bocavirus 1
| Supramolecule | Name: Human bocavirus 1 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 689403 / Sci species name: Human bocavirus 1 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: viral protein 3
| Macromolecule | Name: viral protein 3 / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human bocavirus 1 Human bocavirus 1 |
| Molecular weight | Theoretical: 60.568301 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSDTDIQDQQ PDTVDAPQNT SGGGTGSIGG GKGSGVGIST GGWVGGSHFS DKYVVTKNTR QFITTIQNGH LYKTEAIETT NQSGKSQRC VTTPWTYFNF NQYSCHFSPQ DWQRLTNEYK RFRPKAMQVK IYNLQIKQIL SNGADTTYNN DLTAGVHIFC D GEHAYPNA ...String: MSDTDIQDQQ PDTVDAPQNT SGGGTGSIGG GKGSGVGIST GGWVGGSHFS DKYVVTKNTR QFITTIQNGH LYKTEAIETT NQSGKSQRC VTTPWTYFNF NQYSCHFSPQ DWQRLTNEYK RFRPKAMQVK IYNLQIKQIL SNGADTTYNN DLTAGVHIFC D GEHAYPNA SHPWDEDVMP DLPYKTWKLF QYGYIPIENE LADLDGNAAG GNATEKALLY QMPFFLLENS DHQVLRTGES TE FTFNFDC EWVNNERAYI PPGLMFNPKV PTRRVQYIRQ NGSTAASTGR IQPYSKPTSW MTGPGLLSAQ RVGPQSSDTA PFM VCTNPE GTHINTGAAG FGSGFDPPSG CLAPTNLEYK LQWYQTPEGT GNNGNIIANP SLSMLRDQLL YKGNQTTYNL VGDI WMFPN QVWDRFPITR ENPIWCKKPR ADKHTIMDPF DGSIAMDHPP GTIFIKMAKI PVPTASNADS YLNIYCTGQV SCEIV WEVE RYATKNWRPE RRHTALGMSL GGESNYTPTY HVDPTGAYIQ PTSYDQCMPV KTNINKVL UniProtKB: Minor capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-20 (5k x 3k) / Average electron dose: 63.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)