[English] 日本語
 Yorodumi
Yorodumi- EMDB-8306: CryoEM structure of a beak and feather disease virus-like particl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8306 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of a beak and feather disease virus-like particle encapsidating ssDNA | |||||||||
 Map data Map data | Beak and feather disease virus-like particle containing ssDNA | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Beak and feather disease virus Beak and feather disease virus | |||||||||
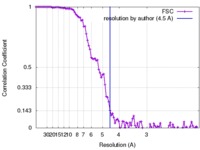
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Hardy JM / Sarker S / Radjainia M / Raidal SR / Forwood JK / Coulibaly F | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2016 Journal: Nat Commun / Year: 2016Title: Structural insights into the assembly and regulation of distinct viral capsid complexes. Authors: Subir Sarker / María C Terrón / Yogesh Khandokar / David Aragão / Joshua M Hardy / Mazdak Radjainia / Manuel Jiménez-Zaragoza / Pedro J de Pablo / Fasséli Coulibaly / Daniel Luque / ...Authors: Subir Sarker / María C Terrón / Yogesh Khandokar / David Aragão / Joshua M Hardy / Mazdak Radjainia / Manuel Jiménez-Zaragoza / Pedro J de Pablo / Fasséli Coulibaly / Daniel Luque / Shane R Raidal / Jade K Forwood /   Abstract: The assembly and regulation of viral capsid proteins into highly ordered macromolecular complexes is essential for viral replication. Here, we utilize crystal structures of the capsid protein from ...The assembly and regulation of viral capsid proteins into highly ordered macromolecular complexes is essential for viral replication. Here, we utilize crystal structures of the capsid protein from the smallest and simplest known viruses capable of autonomously replicating in animal cells, circoviruses, to establish structural and mechanistic insights into capsid morphogenesis and regulation. The beak and feather disease virus, like many circoviruses, encode only two genes: a capsid protein and a replication initiation protein. The capsid protein forms distinct macromolecular assemblies during replication and here we elucidate these structures at high resolution, showing that these complexes reverse the exposure of the N-terminal arginine rich domain responsible for DNA binding and nuclear localization. We show that assembly of these complexes is regulated by single-stranded DNA (ssDNA), and provide a structural basis of capsid assembly around single-stranded DNA, highlighting novel binding interfaces distinct from the highly positively charged N-terminal ARM domain. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8306.map.gz emd_8306.map.gz | 23.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8306-v30.xml emd-8306-v30.xml emd-8306.xml emd-8306.xml | 19.6 KB 19.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8306_fsc.xml emd_8306_fsc.xml | 10.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_8306.png emd_8306.png | 202.2 KB | ||
| Others |  emd_8306_additional.map.gz emd_8306_additional.map.gz | 23.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8306 http://ftp.pdbj.org/pub/emdb/structures/EMD-8306 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8306 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8306 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8306.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8306.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Beak and feather disease virus-like particle containing ssDNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
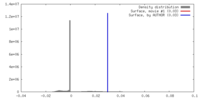
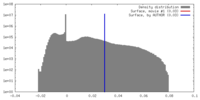
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: The difference map produced by subtracting the crystal...
| File | emd_8306_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The difference map produced by subtracting the crystal structure of the protein component from the cryoEM map | ||||||||||||
| Projections & Slices |
| ||||||||||||
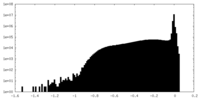
| Density Histograms |
- Sample components
Sample components
-Entire : Beak and feather disease virus
| Entire | Name:  Beak and feather disease virus Beak and feather disease virus |
|---|---|
| Components |
|
-Supramolecule #1: Beak and feather disease virus
| Supramolecule | Name: Beak and feather disease virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: Recombinant protein complex bound to ssDNA in an icosahedral arrangement NCBI-ID: 77856 / Sci species name: Beak and feather disease virus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Cacatua leadbeateri (Major Mitchell's cockatoo) Cacatua leadbeateri (Major Mitchell's cockatoo) |
| Host system | Organism:  |
| Molecular weight | Theoretical: 2.595 MDa |
| Virus shell | Shell ID: 1 / Name: Capsid / Diameter: 170.0 Å / T number (triangulation number): 1 |
-Supramolecule #2: 42mer ssDNA
| Supramolecule | Name: 42mer ssDNA / type: organelle_or_cellular_component / ID: 2 / Parent: 1 / Macromolecule list: #2 Details: The difference map produced by subtracting the crystal structure of the protein component (PDB: 5J37) from the cryoEM map |
|---|---|
| Source (natural) | Organism:  Beak and feather disease virus Beak and feather disease virus |
| Molecular weight | Theoretical: 784 KDa |
-Supramolecule #3: Cap protein
| Supramolecule | Name: Cap protein / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 / Details: Icosahedral capsid |
|---|---|
| Source (natural) | Organism:  Beak and feather disease virus Beak and feather disease virus |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 1.811 MDa |
-Macromolecule #1: Cap Protein
| Macromolecule | Name: Cap Protein / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Beak and feather disease virus Beak and feather disease virus |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHSSG VDLGTENLYF QSNARRRYAR PYRRRHIRRY RRRRRHFRRR RFSTNRIYTL RLTRQFQFKI NKQTTSVGNL IFNADYITFA LDDFLQAVPN PHTLNFEDYR IKLAKMEMRP TGGHYTVQSD GFGHTAVIQD SRITRFKTTA DQTQDPLAPF DGAKKWFVSR ...String: MHHHHHHSSG VDLGTENLYF QSNARRRYAR PYRRRHIRRY RRRRRHFRRR RFSTNRIYTL RLTRQFQFKI NKQTTSVGNL IFNADYITFA LDDFLQAVPN PHTLNFEDYR IKLAKMEMRP TGGHYTVQSD GFGHTAVIQD SRITRFKTTA DQTQDPLAPF DGAKKWFVSR GFKRLLRPKP QITIEDLTTA NQSAALWLNS ARTGWIPLQG GPNSAGTKVR HYGIAFSFPQ PEQTITYVTK LTLYVQFRQF APNNPST |
-Macromolecule #2: 42mer ssDNA
| Macromolecule | Name: 42mer ssDNA / type: dna / ID: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Beak and feather disease virus Beak and feather disease virus |
| Sequence | String: TTATCCACTT CCAATGTTAT TAAGTGCTGG GATTGTTAGG GG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.6 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 3 second blot (-1 force), 2 second drain time. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Sampling interval: 14.0 µm / Digitization - Frames/image: 1-6 / Number grids imaged: 1 / Number real images: 1153 / Average exposure time: 1.0 sec. / Average electron dose: 2.65 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 127000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Fitting, considering the icosahedral symmetries, was done using UCSF Chimera. |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Correlation coefficient |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)