+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6935 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of PCV3 VLP | |||||||||
 Map data Map data | Cryo-EM structure of PCV3 VLP | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | Circovirus capsid protein / Circovirus capsid superfamily / Circovirus capsid protein / viral capsid assembly / T=1 icosahedral viral capsid / symbiont entry into host cell / virion attachment to host cell / Cap Function and homology information Function and homology information | |||||||||
| Biological species |  Porcine circovirus Porcine circovirus | |||||||||
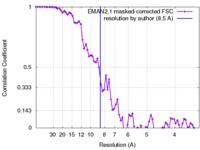
| Method | single particle reconstruction / cryo EM / Resolution: 8.5 Å | |||||||||
 Authors Authors | Mo X / Yuan AY | |||||||||
 Citation Citation |  Journal: to be published Journal: to be publishedTitle: Cryo-EM Structure of Virus-like-particle of Porcine Circovirus type 3 reveals an T=1 symmetry icosahedral particle Authors: Mo X / Yuan AY | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6935.map.gz emd_6935.map.gz | 25.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6935-v30.xml emd-6935-v30.xml emd-6935.xml emd-6935.xml | 12.5 KB 12.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6935_fsc.xml emd_6935_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_6935.png emd_6935.png | 154 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6935 http://ftp.pdbj.org/pub/emdb/structures/EMD-6935 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6935 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6935 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6935.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6935.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of PCV3 VLP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.69 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Porcine circovirus
| Entire | Name:  Porcine circovirus Porcine circovirus |
|---|---|
| Components |
|
-Supramolecule #1: Porcine circovirus
| Supramolecule | Name: Porcine circovirus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Porcine circovirus 3 capsid protein / NCBI-ID: 46221 / Sci species name: Porcine circovirus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host system | Organism: |
| Molecular weight | Experimental: 1.7 MDa |
-Macromolecule #1: PCV3 capsid protein
| Macromolecule | Name: PCV3 capsid protein / type: other / ID: 1 / Classification: other |
|---|---|
| Source (natural) | Organism:  Porcine circovirus Porcine circovirus |
| Sequence | String: MRHRAIFRRR PRPRRRRRHR RRYVRRKLFI RRPTAGTYYT KKYSTMNVIS VGTPQNNKPW HANHFITRLN EWETAISFEY YKILKMKVT LSPVISPAQQ TKTMFGHTAI DLDGAWTTNT WLQDDPYAES STRKVMTSKK KHSRYFTPKP ILAGTTSAHP G QSLFFFSR ...String: MRHRAIFRRR PRPRRRRRHR RRYVRRKLFI RRPTAGTYYT KKYSTMNVIS VGTPQNNKPW HANHFITRLN EWETAISFEY YKILKMKVT LSPVISPAQQ TKTMFGHTAI DLDGAWTTNT WLQDDPYAES STRKVMTSKK KHSRYFTPKP ILAGTTSAHP G QSLFFFSR PTPWLNTYDP TVQWGALLWS IYVPEKTGMT DFYGTKEVWI RYKSVL |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 Component:
Details: 20mM PB(pH 6.5), 500mM NaCl | |||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: FORMVAR / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 101.325 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 5 seconds before pluning. | |||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Details | Preliminary grid screening was performed manually. |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Number grids imaged: 5 / Number real images: 90 / Average exposure time: 1.0 sec. / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.5 µm / Calibrated defocus min: 1.0 µm / Calibrated magnification: 47000 / Illumination mode: OTHER / Imaging mode: DARK FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL / Overall B value: 300 |
|---|
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)