[English] 日本語
 Yorodumi
Yorodumi- EMDB-7519: The H/ACA ribonucleoprotein lobe of the cryo-EM structure of subs... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7519 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The H/ACA ribonucleoprotein lobe of the cryo-EM structure of substrate-bound human telomerase holoenzyme | |||||||||
 Map data Map data | The H/ACA RNP lobe of the human telomerase holoenzyme | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
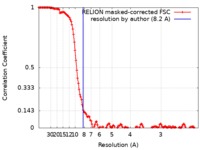
| Method | single particle reconstruction / cryo EM / Resolution: 8.2 Å | |||||||||
 Authors Authors | Nguyen THD / Tam J / Wu RA / Greber BJ / Toso D / Nogales E / Collins K | |||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Cryo-EM structure of substrate-bound human telomerase holoenzyme. Authors: Thi Hoang Duong Nguyen / Jane Tam / Robert A Wu / Basil J Greber / Daniel Toso / Eva Nogales / Kathleen Collins /  Abstract: The enzyme telomerase adds telomeric repeats to chromosome ends to balance the loss of telomeres during genome replication. Telomerase regulation has been implicated in cancer, other human diseases, ...The enzyme telomerase adds telomeric repeats to chromosome ends to balance the loss of telomeres during genome replication. Telomerase regulation has been implicated in cancer, other human diseases, and ageing, but progress towards clinical manipulation of telomerase has been hampered by the lack of structural data. Here we present the cryo-electron microscopy structure of the substrate-bound human telomerase holoenzyme at subnanometre resolution, showing two flexibly RNA-tethered lobes: the catalytic core with telomerase reverse transcriptase (TERT) and conserved motifs of telomerase RNA (hTR), and an H/ACA ribonucleoprotein (RNP). In the catalytic core, RNA encircles TERT, adopting a well-ordered tertiary structure with surprisingly limited protein-RNA interactions. The H/ACA RNP lobe comprises two sets of heterotetrameric H/ACA proteins and one Cajal body protein, TCAB1, representing a pioneering structure of a large eukaryotic family of ribosome and spliceosome biogenesis factors. Our findings provide a structural framework for understanding human telomerase disease mutations and represent an important step towards telomerase-related clinical therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7519.map.gz emd_7519.map.gz | 226.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7519-v30.xml emd-7519-v30.xml emd-7519.xml emd-7519.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_7519_fsc.xml emd_7519_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_7519.png emd_7519.png | 105.4 KB | ||
| Masks |  emd_7519_msk_1.map emd_7519_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7519 http://ftp.pdbj.org/pub/emdb/structures/EMD-7519 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7519 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7519 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7519.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7519.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The H/ACA RNP lobe of the human telomerase holoenzyme | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
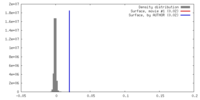
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_7519_msk_1.map emd_7519_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
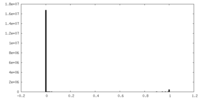
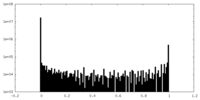
| Projections & Slices |
| ||||||||||||
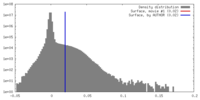
| Density Histograms |
- Sample components
Sample components
-Entire : Human telomerase holoenzyme in complex with DNA substrate
| Entire | Name: Human telomerase holoenzyme in complex with DNA substrate |
|---|---|
| Components |
|
-Supramolecule #1: Human telomerase holoenzyme in complex with DNA substrate
| Supramolecule | Name: Human telomerase holoenzyme in complex with DNA substrate type: complex / ID: 1 / Parent: 0 / Details: Sample was purified from human cells |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Details: unspecified |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 11654 / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)