[English] 日本語
 Yorodumi
Yorodumi- EMDB-6912: Subtomogram average of the budding yeast DASH outer-kinetochore ring. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6912 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of the budding yeast DASH outer-kinetochore ring. | |||||||||
 Map data Map data | Subtomogram average of reconstituted DASH outer-kinetochore rings around microtubules. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM / Resolution: 32.0 Å | |||||||||
 Authors Authors | Ng C / Deng L / Chen C / Lim H / Shi J / Surana U / Gan L | |||||||||
 Citation Citation |  Journal: J Cell Biol / Year: 2019 Journal: J Cell Biol / Year: 2019Title: Electron cryotomography analysis of Dam1C/DASH at the kinetochore-spindle interface in situ. Authors: Cai Tong Ng / Li Deng / Chen Chen / Hong Hwa Lim / Jian Shi / Uttam Surana / Lu Gan /  Abstract: In dividing cells, depolymerizing spindle microtubules move chromosomes by pulling at their kinetochores. While kinetochore subcomplexes have been studied extensively in vitro, little is known about ...In dividing cells, depolymerizing spindle microtubules move chromosomes by pulling at their kinetochores. While kinetochore subcomplexes have been studied extensively in vitro, little is known about their in vivo structure and interactions with microtubules or their response to spindle damage. Here we combine electron cryotomography of serial cryosections with genetic and pharmacological perturbation to study the yeast chromosome segregation machinery in vivo. Each kinetochore microtubule has one (rarely, two) Dam1C/DASH outer kinetochore assemblies. Dam1C/DASH contacts the microtubule walls and does so with its flexible "bridges"; there are no contacts with the protofilaments' curved tips. In metaphase, ∼40% of the Dam1C/DASH assemblies are complete rings; the rest are partial rings. Ring completeness and binding position along the microtubule are sensitive to kinetochore attachment and tension, respectively. Our study and those of others support a model in which each kinetochore must undergo cycles of conformational change to couple microtubule depolymerization to chromosome movement. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6912.map.gz emd_6912.map.gz | 1.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6912-v30.xml emd-6912-v30.xml emd-6912.xml emd-6912.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6912.png emd_6912.png | 87.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6912 http://ftp.pdbj.org/pub/emdb/structures/EMD-6912 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6912 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6912 | HTTPS FTP |
-Related structure data
| Related structure data |  6914C C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10159 (Title: A multi-scale model of the yeast chromosome-segregation system EMPIAR-10159 (Title: A multi-scale model of the yeast chromosome-segregation systemData size: 95.4 Data #1: Tilt series, tomograms, and models of cryosections of mitotic budding yeast and reconstituted DASH rings around microtubules [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6912.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6912.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of reconstituted DASH outer-kinetochore rings around microtubules. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.61 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
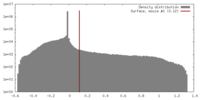
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Budding yeast DASH outer-kinetochore ring complex
| Entire | Name: Budding yeast DASH outer-kinetochore ring complex |
|---|---|
| Components |
|
-Supramolecule #1: Budding yeast DASH outer-kinetochore ring complex
| Supramolecule | Name: Budding yeast DASH outer-kinetochore ring complex / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 3.4 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 6.8 Details: 20 mM sodium phosphate pH 6.8, 150 mM NaCl, 1 mM EDTA |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE |
| Details | DASH complexes mixed with Taxol-stabilized microtubules |
| Sectioning | Other: NO SECTIONING |
| Fiducial marker | Manufacturer: BBI / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Sampling interval: 14.0 µm / Number grids imaged: 1 / Number real images: 67 / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 30400 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 14.0 µm / Nominal defocus min: 8.0 µm / Nominal magnification: 18000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 32.0 Å / Resolution method: FSC 0.143 CUT-OFF Software:
Number images used: 66 | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Software - Name: eTomo (ver. 4.9) / Details: CTF compensation done in Etomo |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)