[English] 日本語
 Yorodumi
Yorodumi- EMDB-6703: Prefusion structure of SARS-CoV spike glycoprotein, conformation 1 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6703 | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Prefusion structure of SARS-CoV spike glycoprotein, conformation 1 | ||||||||||||||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||||||||
 Keywords Keywords | SARS-CoV / spike glycoprotein / prefusion / single particle / VIRAL PROTEIN | ||||||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / Attachment and Entry / SARS-CoV-1 activates/modulates innate immune responses / symbiont-mediated-mediated suppression of host tetherin activity / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell ...Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / Attachment and Entry / SARS-CoV-1 activates/modulates innate immune responses / symbiont-mediated-mediated suppression of host tetherin activity / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane / membrane / identical protein binding Similarity search - Function | ||||||||||||||||||||||||||||||||||||
| Biological species |  SARS coronavirus BJ01 SARS coronavirus BJ01 | ||||||||||||||||||||||||||||||||||||
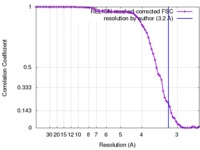
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | ||||||||||||||||||||||||||||||||||||
 Authors Authors | Yuan Y / Cao D | ||||||||||||||||||||||||||||||||||||
| Funding support |  China, 11 items China, 11 items
| ||||||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Authors: Yuan Yuan / Duanfang Cao / Yanfang Zhang / Jun Ma / Jianxun Qi / Qihui Wang / Guangwen Lu / Ying Wu / Jinghua Yan / Yi Shi / Xinzheng Zhang / George F Gao /  Abstract: The envelope spike (S) proteins of MERS-CoV and SARS-CoV determine the virus host tropism and entry into host cells, and constitute a promising target for the development of prophylactics and ...The envelope spike (S) proteins of MERS-CoV and SARS-CoV determine the virus host tropism and entry into host cells, and constitute a promising target for the development of prophylactics and therapeutics. Here, we present high-resolution structures of the trimeric MERS-CoV and SARS-CoV S proteins in its pre-fusion conformation by single particle cryo-electron microscopy. The overall structures resemble that from other coronaviruses including HKU1, MHV and NL63 reported recently, with the exception of the receptor binding domain (RBD). We captured two states of the RBD with receptor binding region either buried (lying state) or exposed (standing state), demonstrating an inherently flexible RBD readily recognized by the receptor. Further sequence conservation analysis of six human-infecting coronaviruses revealed that the fusion peptide, HR1 region and the central helix are potential targets for eliciting broadly neutralizing antibodies. | ||||||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6703.map.gz emd_6703.map.gz | 28.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6703-v30.xml emd-6703-v30.xml emd-6703.xml emd-6703.xml | 13.9 KB 13.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6703_fsc.xml emd_6703_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_6703.png emd_6703.png | 129 KB | ||
| Filedesc metadata |  emd-6703.cif.gz emd-6703.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6703 http://ftp.pdbj.org/pub/emdb/structures/EMD-6703 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6703 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6703 | HTTPS FTP |
-Related structure data
| Related structure data |  5x58MC  6704C  6705C  6706C  6707C  5x4rC  5x4sC  5x59C  5x5bC  5x5cC  5x5fC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6703.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6703.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SARS-CoV spike trimer
| Entire | Name: SARS-CoV spike trimer |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV spike trimer
| Supramolecule | Name: SARS-CoV spike trimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  SARS coronavirus BJ01 / Strain: BJ01 SARS coronavirus BJ01 / Strain: BJ01 |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  SARS coronavirus BJ01 / Strain: BJ01 SARS coronavirus BJ01 / Strain: BJ01 |
| Molecular weight | Theoretical: 136.825891 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SDLDRCTTFD DVQAPNYTQH TSSMRGVYYP DEIFRSDTLY LTQDLFLPFY SNVTGFHTIN HTFDNPVIPF KDGIYFAATE KSNVVRGWV FGSTMNNKSQ SVIIINNSTN VVIRACNFEL CDNPFFAVSK PMGTQTHTMI FDNAFNCTFE YISDAFSLDV S EKSGNFKH ...String: SDLDRCTTFD DVQAPNYTQH TSSMRGVYYP DEIFRSDTLY LTQDLFLPFY SNVTGFHTIN HTFDNPVIPF KDGIYFAATE KSNVVRGWV FGSTMNNKSQ SVIIINNSTN VVIRACNFEL CDNPFFAVSK PMGTQTHTMI FDNAFNCTFE YISDAFSLDV S EKSGNFKH LREFVFKNKD GFLYVYKGYQ PIDVVRDLPS GFNTLKPIFK LPLGINITNF RAILTAFSPA QDTWGTSAAA YF VGYLKPT TFMLKYDENG TITDAVDCSQ NPLAELKCSV KSFEIDKGIY QTSNFRVVPS GDVVRFPNIT NLCPFGEVFN ATK FPSVYA WERKKISNCV ADYSVLYNST FFSTFKCYGV SATKLNDLCF SNVYADSFVV KGDDVRQIAP GQTGVIADYN YKLP DDFMG CVLAWNTRNI DATSTGNYNY KYRYLRHGKL RPFERDISNV PFSPDGKPCT PPALNCYWPL NDYGFYTTTG IGYQP YRVV VLSFELLNAP ATVCGPKLST DLIKNQCVNF NFNGLTGTGV LTPSSKRFQP FQQFGRDVSD FTDSVRDPKT SEILDI SPC SFGGVSVITP GTNASSEVAV LYQDVNCTDV STAIHADQLT PAWRIYSTGN NVFQTQAGCL IGAEHVDTSY ECDIPIG AG ICASYHTVSL LRSTSQKSIV AYTMSLGADS SIAYSNNTIA IPTNFSISIT TEVMPVSMAK TSVDCNMYIC GDSTECAN L LLQYGSFCTQ LNRALSGIAA EQDRNTREVF AQVKQMYKTP TLKYFGGFNF SQILPDPLKP TKRSFIEDLL FNKVTLADA GFMKQYGECL GDINARDLIC AQKFNGLTVL PPLLTDDMIA AYTAALVSGT ATAGWTFGAG AALQIPFAMQ MAYRFNGIGV TQNVLYENQ KQIANQFNKA ISQIQESLTT TSTALGKLQD VVNQNAQALN TLVKQLSSNF GAISSVLNDI LSRLDKVEAE V QIDRLITG RLQSLQTYVT QQLIRAAEIR ASANLAATKM SECVLGQSKR VDFCGKGYHL MSFPQAAPHG VVFLHVTYVP SQ ERNFTTA PAICHEGKAY FPREGVFVFN GTSWFITQRN FFSPQIITTD NTFVSGNCDV VIGIINNTVY DPLQPELDSF KEE LDKYFK NHTSPDVDLG DISGINASVV NIQKEIDRLN EVAKNLNESL IDLQELGKYE QYIKIKRMKQ IEDKIEEIES KQKK IENEI ARIKKIKLVP RGSLEWSHPQ FEK |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 42 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 8.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Refinement | Protocol: RIGID BODY FIT | ||||||
| Output model |  PDB-5x58: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)