[English] 日本語
 Yorodumi
Yorodumi- EMDB-6697: Cryo-EM structure of the 90S small subunit pre-ribosome (Noc4-TAP) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6697 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the 90S small subunit pre-ribosome (Noc4-TAP) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of ribosomal protein gene transcription by RNA polymerase II / box H/ACA snoRNA binding / RNA fragment catabolic process / rRNA small subunit pseudouridine methyltransferase Nep1 / CURI complex / UTP-C complex / rRNA 2'-O-methylation / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / t-UTP complex / Pwp2p-containing subcomplex of 90S preribosome ...regulation of ribosomal protein gene transcription by RNA polymerase II / box H/ACA snoRNA binding / RNA fragment catabolic process / rRNA small subunit pseudouridine methyltransferase Nep1 / CURI complex / UTP-C complex / rRNA 2'-O-methylation / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / t-UTP complex / Pwp2p-containing subcomplex of 90S preribosome / Mpp10 complex / nuclear microtubule / rRNA (pseudouridine) methyltransferase activity / box C/D sno(s)RNA binding / snoRNA guided rRNA 2'-O-methylation / rRNA modification / histone H2AQ104 methyltransferase activity / septum digestion after cytokinesis / snRNA binding / box C/D sno(s)RNA 3'-end processing / rRNA methyltransferase activity / endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / regulation of rRNA processing / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, LSU-rRNA,5S) / regulation of transcription by RNA polymerase I / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / RNA folding chaperone / box C/D methylation guide snoRNP complex / rDNA heterochromatin / U4/U6 snRNP / tRNA export from nucleus / single-stranded telomeric DNA binding / rRNA primary transcript binding / O-methyltransferase activity / sno(s)RNA-containing ribonucleoprotein complex / translational readthrough / : / rRNA base methylation / protein localization to nucleolus / SUMOylation of RNA binding proteins / rRNA methylation / U4 snRNA binding / mTORC1-mediated signalling / Protein hydroxylation / 90S preribosome assembly / U4 snRNP / U3 snoRNA binding / positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / Formation of the ternary complex, and subsequently, the 43S complex / poly(U) RNA binding / Translation initiation complex formation / Ribosomal scanning and start codon recognition / poly(A)+ mRNA export from nucleus / snoRNA binding / preribosome, small subunit precursor / precatalytic spliceosome / Major pathway of rRNA processing in the nucleolus and cytosol / establishment of cell polarity / SRP-dependent cotranslational protein targeting to membrane / positive regulation of transcription by RNA polymerase I / GTP hydrolysis and joining of the 60S ribosomal subunit / nucleolar large rRNA transcription by RNA polymerase I / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / spliceosomal complex assembly / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / 90S preribosome / Ub-specific processing proteases / ribosomal subunit export from nucleus / proteasome assembly / U4/U6 x U5 tri-snRNP complex / regulation of translational fidelity / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal small subunit export from nucleus / RNA endonuclease activity / nuclear periphery / Transferases; Transferring one-carbon groups; Methyltransferases / ribosome assembly / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / spliceosomal complex / maturation of SSU-rRNA / translational initiation / small-subunit processome / mRNA splicing, via spliceosome / maintenance of translational fidelity / modification-dependent protein catabolic process / enzyme activator activity / cytoplasmic stress granule / protein tag activity / rRNA processing / : / ribosomal small subunit assembly / ribosome biogenesis / ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / cytoplasmic translation Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.7 Å | |||||||||
 Authors Authors | Ye K / Zhu X / Sun Q | |||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: Molecular architecture of the 90S small subunit pre-ribosome. Authors: Qi Sun / Xing Zhu / Jia Qi / Weidong An / Pengfei Lan / Dan Tan / Rongchang Chen / Bing Wang / Sanduo Zheng / Cheng Zhang / Xining Chen / Wei Zhang / Jing Chen / Meng-Qiu Dong / Keqiong Ye /  Abstract: Eukaryotic small ribosomal subunits are first assembled into 90S pre-ribosomes. The complete 90S is a gigantic complex with a molecular mass of approximately five megadaltons. Here, we report the ...Eukaryotic small ribosomal subunits are first assembled into 90S pre-ribosomes. The complete 90S is a gigantic complex with a molecular mass of approximately five megadaltons. Here, we report the nearly complete architecture of 90S determined from three cryo-electron microscopy single particle reconstructions at 4.5 to 8.7 angstrom resolution. The majority of the density maps were modeled and assigned to specific RNA and protein components. The nascent ribosome is assembled into isolated native-like substructures that are stabilized by abundant assembly factors. The 5' external transcribed spacer and U3 snoRNA nucleate a large subcomplex that scaffolds the nascent ribosome. U3 binds four sites of pre-rRNA, including a novel site on helix 27 but not the 3' side of the central pseudoknot, and crucially organizes the 90S structure. The 90S model provides significant insight into the principle of small subunit assembly and the function of assembly factors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6697.map.gz emd_6697.map.gz | 390 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6697-v30.xml emd-6697-v30.xml emd-6697.xml emd-6697.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6697_fsc.xml emd_6697_fsc.xml | 16 KB | Display |  FSC data file FSC data file |
| Images |  emd_6697.png emd_6697.png | 84.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6697 http://ftp.pdbj.org/pub/emdb/structures/EMD-6697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6697 | HTTPS FTP |
-Related structure data
| Related structure data |  6695C  6696C  5wwnC  5wwoC  5wxlC  5wxmC  5wy3C  5wyjC  5wykC  5wylC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6697.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6697.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.42 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
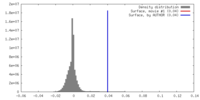
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 90S small subunit pre-ribosome (Noc4-TAP)
| Entire | Name: 90S small subunit pre-ribosome (Noc4-TAP) |
|---|---|
| Components |
|
-Supramolecule #1: 90S small subunit pre-ribosome (Noc4-TAP)
| Supramolecule | Name: 90S small subunit pre-ribosome (Noc4-TAP) / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 5.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 101.325 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | OD280=2.0 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4000 pixel / Digitization - Dimensions - Height: 4000 pixel / Number grids imaged: 1 / Number real images: 1769 / Average exposure time: 1.6 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 98592 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)