[English] 日本語
 Yorodumi
Yorodumi- EMDB-6493: Electron cryo-microscopy of bacteriophage EL chaperonin in the AD... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6493 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron cryo-microscopy of bacteriophage EL chaperonin in the ADP-bound conformation | |||||||||
 Map data Map data | Reconstruction of the bacteriophage phi EL ADP-bound chaperonin conformation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | chaperonin / phi-EL / protein folding / ADP conformation | |||||||||
| Function / homology |  Function and homology information Function and homology informationATP-dependent protein folding chaperone / protein refolding / ATP binding / metal ion binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Pseudomonas phage EL (virus) Pseudomonas phage EL (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 11.7 Å | |||||||||
 Authors Authors | Molugu SK / Hildenbrand ZL / Morgan DG / Sherman MB / He L / Georgopoulos C / Sernova NV / Kurochkina LP / Mesyanzhinov VV / Miroshnikov KA / Bernal RA | |||||||||
 Citation Citation |  Journal: Structure / Year: 2016 Journal: Structure / Year: 2016Title: Ring Separation Highlights the Protein-Folding Mechanism Used by the Phage EL-Encoded Chaperonin. Authors: Sudheer K Molugu / Zacariah L Hildenbrand / David Gene Morgan / Michael B Sherman / Lilin He / Costa Georgopoulos / Natalia V Sernova / Lidia P Kurochkina / Vadim V Mesyanzhinov / Konstantin ...Authors: Sudheer K Molugu / Zacariah L Hildenbrand / David Gene Morgan / Michael B Sherman / Lilin He / Costa Georgopoulos / Natalia V Sernova / Lidia P Kurochkina / Vadim V Mesyanzhinov / Konstantin A Miroshnikov / Ricardo A Bernal /   Abstract: Chaperonins are ubiquitous, ATP-dependent protein-folding molecular machines that are essential for all forms of life. Bacteriophage φEL encodes its own chaperonin to presumably fold exceedingly ...Chaperonins are ubiquitous, ATP-dependent protein-folding molecular machines that are essential for all forms of life. Bacteriophage φEL encodes its own chaperonin to presumably fold exceedingly large viral proteins via profoundly different nucleotide-binding conformations. Our structural investigations indicate that ATP likely binds to both rings simultaneously and that a misfolded substrate acts as the trigger for ATP hydrolysis. More importantly, the φEL complex dissociates into two single rings resulting from an evolutionarily altered residue in the highly conserved ATP-binding pocket. Conformational changes also more than double the volume of the single-ring internal chamber such that larger viral proteins are accommodated. This is illustrated by the fact that φEL is capable of folding β-galactosidase, a 116-kDa protein. Collectively, the architecture and protein-folding mechanism of the φEL chaperonin are significantly different from those observed in group I and II chaperonins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6493.map.gz emd_6493.map.gz | 14.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6493-v30.xml emd-6493-v30.xml emd-6493.xml emd-6493.xml | 10.4 KB 10.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6493.jpg emd_6493.jpg | 96.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6493 http://ftp.pdbj.org/pub/emdb/structures/EMD-6493 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6493 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6493 | HTTPS FTP |
-Validation report
| Summary document |  emd_6493_validation.pdf.gz emd_6493_validation.pdf.gz | 79.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6493_full_validation.pdf.gz emd_6493_full_validation.pdf.gz | 78.4 KB | Display | |
| Data in XML |  emd_6493_validation.xml.gz emd_6493_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6493 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6493 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6493 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6493 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6493.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6493.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of the bacteriophage phi EL ADP-bound chaperonin conformation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.17 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
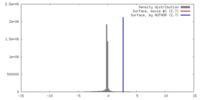
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriphage phi-EL encoded chaperonin in the ADP-bound conformation
| Entire | Name: Bacteriphage phi-EL encoded chaperonin in the ADP-bound conformation |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriphage phi-EL encoded chaperonin in the ADP-bound conformation
| Supramolecule | Name: Bacteriphage phi-EL encoded chaperonin in the ADP-bound conformation type: sample / ID: 1000 / Oligomeric state: homoheptamer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 431.7677 KDa |
-Macromolecule #1: phi-EL chaperonin
| Macromolecule | Name: phi-EL chaperonin / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Oligomeric state: heptamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage EL (virus) / synonym: Bacteriophage EL Pseudomonas phage EL (virus) / synonym: Bacteriophage EL |
| Molecular weight | Theoretical: 61.6811 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Putative GroEL-like chaperonine protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 50 mM HEPES, pH 7.5, 150 mM NaCl, 2 mM ADP, 2 mM EDTA |
| Grid | Details: Quantifoil R2/2 grids glow-discharged in air for 1 minute |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 100 K / Instrument: HOMEMADE PLUNGER / Method: Blot for 2-3 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FS |
|---|---|
| Temperature | Min: 88 K / Max: 103 K / Average: 100 K |
| Date | Aug 1, 2009 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Sampling interval: 12 µm / Number real images: 50 / Average electron dose: 10 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 69000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 11.7 Å / Resolution method: OTHER / Software - Name: EMAN, EMAN2, RELION / Number images used: 18778 |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)