+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6448 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of F-actin | |||||||||
 Map data Map data | Control reconstruction of F-actin alone | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | actin / cell migration / adhesion / mechanosensation / cytoskeleton | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoskeletal motor activator activity / myosin heavy chain binding / tropomyosin binding / actin filament bundle / troponin I binding / filamentous actin / mesenchyme migration / skeletal muscle myofibril / actin filament bundle assembly / striated muscle thin filament ...cytoskeletal motor activator activity / myosin heavy chain binding / tropomyosin binding / actin filament bundle / troponin I binding / filamentous actin / mesenchyme migration / skeletal muscle myofibril / actin filament bundle assembly / striated muscle thin filament / skeletal muscle thin filament assembly / actin monomer binding / skeletal muscle fiber development / stress fiber / titin binding / actin filament polymerization / filopodium / actin filament / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / calcium-dependent protein binding / lamellipodium / cell body / protein domain specific binding / hydrolase activity / calcium ion binding / positive regulation of gene expression / magnesium ion binding / ATP binding / identical protein binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 7.6 Å | |||||||||
 Authors Authors | Kim LY / Thompson PM / Lee HT / Pershad M / Campbell SL / Alushin GM | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2016 Journal: J Mol Biol / Year: 2016Title: The Structural Basis of Actin Organization by Vinculin and Metavinculin. Authors: Laura Y Kim / Peter M Thompson / Hyunna T Lee / Mihir Pershad / Sharon L Campbell / Gregory M Alushin /  Abstract: Vinculin is an essential adhesion protein that links membrane-bound integrin and cadherin receptors through their intracellular binding partners to filamentous actin, facilitating mechanotransduction. ...Vinculin is an essential adhesion protein that links membrane-bound integrin and cadherin receptors through their intracellular binding partners to filamentous actin, facilitating mechanotransduction. Here we present an 8.5-Å-resolution cryo-electron microscopy reconstruction and pseudo-atomic model of the vinculin tail (Vt) domain bound to F-actin. Upon actin engagement, the N-terminal "strap" and helix 1 are displaced from the Vt helical bundle to mediate actin bundling. We find that an analogous conformational change also occurs in the H1' helix of the tail domain of metavinculin (MVt) upon actin binding, a muscle-specific splice isoform that suppresses actin bundling by Vt. These data support a model in which metavinculin tunes the actin bundling activity of vinculin in a tissue-specific manner, providing a mechanistic framework for understanding metavinculin mutations associated with hereditary cardiomyopathies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6448.map.gz emd_6448.map.gz | 25.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6448-v30.xml emd-6448-v30.xml emd-6448.xml emd-6448.xml | 10.7 KB 10.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6448.png emd_6448.png | 168 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6448 http://ftp.pdbj.org/pub/emdb/structures/EMD-6448 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6448 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6448 | HTTPS FTP |
-Related structure data
| Related structure data |  3jbjMC  6446C  6447C  6449C  6450C  6451C  3jbiC  3jbkC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6448.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6448.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Control reconstruction of F-actin alone | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
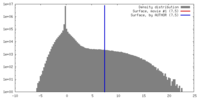
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : F-actin
| Entire | Name: F-actin |
|---|---|
| Components |
|
-Supramolecule #1000: F-actin
| Supramolecule | Name: F-actin / type: sample / ID: 1000 / Details: Helical filament of F-actin / Oligomeric state: Helical / Number unique components: 1 |
|---|
-Macromolecule #1: skeletal muscle actin
| Macromolecule | Name: skeletal muscle actin / type: protein_or_peptide / ID: 1 / Name.synonym: actin / Oligomeric state: helical filament / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.8 KDa |
| Sequence | UniProtKB: Actin, alpha skeletal muscle |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.0125 mg/mL |
|---|---|
| Buffer | pH: 7 / Details: 50 mM KCl, 1 mM MgCl2, 1 mM EGTA, 10 mM imidazole |
| Grid | Details: 200 mesh 1.2 / 1.3 C-flat |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: LEICA EM GP Method: 3 microliters of 0.3 micromolar actin was applied to the grid and incubated for 60 seconds at 25 degrees C. The grid was blotted for 2 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 20 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000 times magnification. |
| Date | May 13, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 1257 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 137615 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 100000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | Single-model IHRSR was performed with EMAN2 / SPARX and final reconstruction with FREALIGN (fixed helical parameters). |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 27.80 Å Applied symmetry - Helical parameters - Δ&Phi: 166.67 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.6 Å / Resolution method: OTHER / Software - Name: CTFFIND3, EMAN2/SPARX, FREALIGN |
| CTF correction | Details: FREALIGN (per segment) |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)