+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22219 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of CARD8-CARD filament | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Filament / inflammasome / signaling / UPA / FIIND / CARD / NLRP1 / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationCARD8 inflammasome complex assembly / CARD8 inflammasome complex / NACHT domain binding / Formation of apoptosome / CARD domain binding / self proteolysis / NLRP3 inflammasome complex / negative regulation of lipopolysaccharide-mediated signaling pathway / negative regulation of NLRP3 inflammasome complex assembly / cysteine-type endopeptidase activator activity ...CARD8 inflammasome complex assembly / CARD8 inflammasome complex / NACHT domain binding / Formation of apoptosome / CARD domain binding / self proteolysis / NLRP3 inflammasome complex / negative regulation of lipopolysaccharide-mediated signaling pathway / negative regulation of NLRP3 inflammasome complex assembly / cysteine-type endopeptidase activator activity / Regulation of the apoptosome activity / Hydrolases; Acting on peptide bonds (peptidases) / negative regulation of interleukin-1 beta production / pattern recognition receptor activity / negative regulation of tumor necrosis factor-mediated signaling pathway / activation of innate immune response / intrinsic apoptotic signaling pathway / antiviral innate immune response / positive regulation of interleukin-1 beta production / negative regulation of canonical NF-kappaB signal transduction / molecular condensate scaffold activity / protein destabilization / positive regulation of inflammatory response / peptidase activity / regulation of apoptotic process / defense response to virus / inflammatory response / protein homodimerization activity / protein-containing complex / nucleoplasm / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
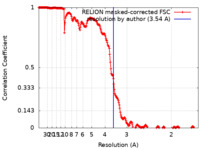
| Method | helical reconstruction / cryo EM / Resolution: 3.54 Å | |||||||||
 Authors Authors | Hollingsworth LR / David L | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Mechanism of filament formation in UPA-promoted CARD8 and NLRP1 inflammasomes. Authors: L Robert Hollingsworth / Liron David / Yang Li / Andrew R Griswold / Jianbin Ruan / Humayun Sharif / Pietro Fontana / Elizabeth L Orth-He / Tian-Min Fu / Daniel A Bachovchin / Hao Wu /  Abstract: NLRP1 and CARD8 are related cytosolic sensors that upon activation form supramolecular signalling complexes known as canonical inflammasomes, resulting in caspase-1 activation, cytokine maturation ...NLRP1 and CARD8 are related cytosolic sensors that upon activation form supramolecular signalling complexes known as canonical inflammasomes, resulting in caspase-1 activation, cytokine maturation and/or pyroptotic cell death. NLRP1 and CARD8 use their C-terminal (CT) fragments containing a caspase recruitment domain (CARD) and the UPA (conserved in UNC5, PIDD, and ankyrins) subdomain for self-oligomerization, which in turn form the platform to recruit the inflammasome adaptor ASC (apoptosis-associated speck-like protein containing a CARD) or caspase-1, respectively. Here, we report cryo-EM structures of NLRP1-CT and CARD8-CT assemblies, in which the respective CARDs form central helical filaments that are promoted by oligomerized, but flexibly linked, UPAs surrounding the filaments. Through biochemical and cellular approaches, we demonstrate that the UPA itself reduces the threshold needed for NLRP1-CT and CARD8-CT filament formation and signalling. Structural analyses provide insights on the mode of ASC recruitment by NLRP1-CT and the contrasting direct recruitment of caspase-1 by CARD8-CT. We also discover that subunits in the central NLRP1 filament dimerize with additional exterior CARDs, which roughly doubles its thickness and is unique among all known CARD filaments. Finally, we engineer and determine the structure of an ASC-caspase-1 octamer, which suggests that ASC uses opposing surfaces for NLRP1, versus caspase-1, recruitment. Together these structures capture the architecture and specificity of the active NLRP1 and CARD8 inflammasomes in addition to key heteromeric CARD-CARD interactions governing inflammasome signalling. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22219.map.gz emd_22219.map.gz | 925.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22219-v30.xml emd-22219-v30.xml emd-22219.xml emd-22219.xml | 13.1 KB 13.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22219_fsc.xml emd_22219_fsc.xml | 22.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_22219.png emd_22219.png | 139.8 KB | ||
| Filedesc metadata |  emd-22219.cif.gz emd-22219.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22219 http://ftp.pdbj.org/pub/emdb/structures/EMD-22219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22219 | HTTPS FTP |
-Related structure data
| Related structure data |  6xkjMC  6xkkC  7keuC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10567 (Title: CARD8-CT filament / Data size: 1.2 TB EMPIAR-10567 (Title: CARD8-CT filament / Data size: 1.2 TBData #1: Unaligned multi-frame micrographs for the CARD8 CT filament [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22219.map.gz / Format: CCP4 / Size: 1000 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22219.map.gz / Format: CCP4 / Size: 1000 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8315 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
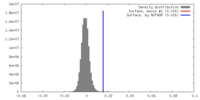
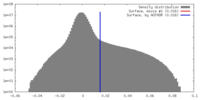
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : CARD8-CARD filament
| Entire | Name: CARD8-CARD filament |
|---|---|
| Components |
|
-Supramolecule #1: CARD8-CARD filament
| Supramolecule | Name: CARD8-CARD filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Caspase recruitment domain-containing protein 8
| Macromolecule | Name: Caspase recruitment domain-containing protein 8 / type: protein_or_peptide / ID: 1 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.103373 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AAFVKENHRQ LQARMGDLKG VLDDLQDNEV LTENEKELVE QEKTRQSKNE ALLSMVEKKG DLALDVLFRS ISERDPYLVS YLRQQNL UniProtKB: Caspase recruitment domain-containing protein 8 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1208 / Average exposure time: 1.25 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -2.2 µm / Nominal defocus min: -0.8 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)