+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6410 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of PhnGHIJK complex from Escherichia coli | |||||||||
 Map data Map data | PhnGHIJK complex from Escherichia coli | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PhnGHIJK | |||||||||
| Biological species |  | |||||||||
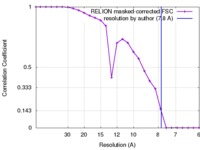
| Method | single particle reconstruction / cryo EM / Resolution: 7.8 Å | |||||||||
 Authors Authors | Yang K / Ren Z / Raushel FM / Zhang J | |||||||||
 Citation Citation |  Journal: Structure / Year: 2016 Journal: Structure / Year: 2016Title: Structures of the Carbon-Phosphorus Lyase Complex Reveal the Binding Mode of the NBD-like PhnK. Authors: Kailu Yang / Zhongjie Ren / Frank M Raushel / Junjie Zhang /  Abstract: The carbon-phosphorus (C-P) lyase complex is essential for the metabolism of unactivated phosphonates to phosphate in bacteria. Using single-particle cryo-electron microscopy, we determined two ...The carbon-phosphorus (C-P) lyase complex is essential for the metabolism of unactivated phosphonates to phosphate in bacteria. Using single-particle cryo-electron microscopy, we determined two structures of the C-P lyase core complex PhnG2H2I2J2, with or without PhnK. PhnG2H2I2J2 is a two-fold symmetric hetero-octamer. Its two PhnJ subunits provide two identical binding sites for PhnK. Only one PhnK binds to PhnG2H2I2J2 due to steric hindrance. PhnK is homologous to the nucleotide-binding domain (NBD) of ATP-binding cassette transporters. The α helices 3 and 4 of PhnK bind to α helix 6 and a loop (residues 227-230) of PhnJ, in a different mode from the binding of NBDs to their transmembrane partners. Moreover, binding of PhnK exposes the active site residue, Gly32 of PhnJ, located near the interface between PhnJ and PhnH. This structural information provides a basis for further deciphering of the reaction mechanism of the C-P lyase. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6410.map.gz emd_6410.map.gz | 154.7 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6410-v30.xml emd-6410-v30.xml emd-6410.xml emd-6410.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6410_fsc.xml emd_6410_fsc.xml | 2.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_6410.png emd_6410.png | 98.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6410 http://ftp.pdbj.org/pub/emdb/structures/EMD-6410 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6410 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6410 | HTTPS FTP |
-Validation report
| Summary document |  emd_6410_validation.pdf.gz emd_6410_validation.pdf.gz | 78.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6410_full_validation.pdf.gz emd_6410_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_6410_validation.xml.gz emd_6410_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6410 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6410 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6410 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6410 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6410.map.gz / Format: CCP4 / Size: 825.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6410.map.gz / Format: CCP4 / Size: 825.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PhnGHIJK complex from Escherichia coli | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
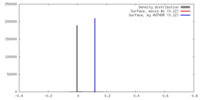
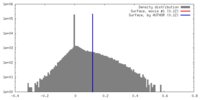
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PhnGHIJK complex from Escherichia coli
| Entire | Name: PhnGHIJK complex from Escherichia coli |
|---|---|
| Components |
|
-Supramolecule #1000: PhnGHIJK complex from Escherichia coli
| Supramolecule | Name: PhnGHIJK complex from Escherichia coli / type: sample / ID: 1000 Oligomeric state: Two PhnG, two PhnH, two PhnI, two PhnJ, one PhnK Number unique components: 5 |
|---|---|
| Molecular weight | Theoretical: 248 KDa |
-Macromolecule #1: PhnG
| Macromolecule | Name: PhnG / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.7 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: PhnH
| Macromolecule | Name: PhnH / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 20.9 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #3: PhnI
| Macromolecule | Name: PhnI / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.9 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #4: PhnJ
| Macromolecule | Name: PhnJ / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 31.7 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #5: PhnK
| Macromolecule | Name: PhnK / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.8 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 8.5 / Details: 50 mM HEPES, 150 mM NaCl, 2 mM TCEP |
| Grid | Details: C-Flat 200 mesh 1.2/1.3 holey carbon grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 91 K / Instrument: FEI VITROBOT MARK III / Method: Blot for 5 seconds before plunging |
- Electron microscopy #1
Electron microscopy #1
| Microscopy ID | 1 |
|---|---|
| Microscope | FEI TECNAI F20 |
| Temperature | Min: 77 K / Max: 105 K / Average: 100 K |
| Details | Counting mode with a dose rate of 10 e/(pixel^2*s), 0.2 s/frame |
| Date | Nov 14, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 967 / Average electron dose: 22 e/Å2 Details: Every image is the average of twenty-five or fifty frames recorded by the direct electron detector. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 33333 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 25000 |
| Sample stage | Specimen holder: Nitrogen-cooled / Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Electron microscopy #2
Electron microscopy #2
| Microscopy ID | 2 |
|---|---|
| Microscope | FEI TECNAI F20 |
| Temperature | Min: 77 K / Max: 105 K / Average: 100 K |
| Details | Counting mode with a dose rate of 10 e/(pixel^2*s), 0.2 s/frame |
| Date | Nov 29, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 967 / Average electron dose: 44 e/Å2 Details: Every image is the average of twenty-five or fifty frames recorded by the direct electron detector. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 33333 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 25000 |
| Sample stage | Specimen holder: Nitrogen-cooled / Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Electron microscopy #3
Electron microscopy #3
| Microscopy ID | 3 |
|---|---|
| Microscope | FEI TECNAI F20 |
| Temperature | Min: 77 K / Max: 105 K / Average: 100 K |
| Details | Counting mode with a dose rate of 10 e/(pixel^2*s), 0.2 s/frame |
| Date | Dec 14, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 967 / Average electron dose: 44 e/Å2 Details: Every image is the average of twenty-five or fifty frames recorded by the direct electron detector. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 33333 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 25000 |
| Sample stage | Specimen holder: Nitrogen-cooled / Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name:  Chimera Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)