[English] 日本語
 Yorodumi
Yorodumi- EMDB-6241: The Structure of a Biologically Active Estrogen Receptor-Coactiva... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6241 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Structure of a Biologically Active Estrogen Receptor-Coactivator Complex on DNA | |||||||||
 Map data Map data | ERa/SRC-3/p300 complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Estrogen receptor | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 25.0 Å | |||||||||
 Authors Authors | Yi P / Wang Z / Feng Q / Pintilie GD / Foulds CE / Lanz RB / Ludtke SJ / Schmid MF / Chiu W / O'Malley BW | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2015 Journal: Mol Cell / Year: 2015Title: Structure of a biologically active estrogen receptor-coactivator complex on DNA. Authors: Ping Yi / Zhao Wang / Qin Feng / Grigore D Pintilie / Charles E Foulds / Rainer B Lanz / Steven J Ludtke / Michael F Schmid / Wah Chiu / Bert W O'Malley /  Abstract: Estrogen receptor (ER/ESR1) is a transcription factor critical for development, reproduction, metabolism, and cancer. ER function hinges on its ability to recruit primary and secondary coactivators, ...Estrogen receptor (ER/ESR1) is a transcription factor critical for development, reproduction, metabolism, and cancer. ER function hinges on its ability to recruit primary and secondary coactivators, yet structural information on the full-length receptor-coactivator complex to complement preexisting and sometimes controversial biochemical information is lacking. Here, we use cryoelectron microscopy (cryo-EM) to determine the quaternary structure of an active complex of DNA-bound ERα, steroid receptor coactivator 3 (SRC-3/NCOA3), and a secondary coactivator (p300/EP300). Our structural model suggests the following assembly mechanism for the complex: each of the two ligand-bound ERα monomers independently recruits one SRC-3 protein via the transactivation domain of ERα; the two SRC-3s in turn bind to different regions of one p300 protein through multiple contacts. We also present structural evidence for the location of activation function 1 (AF-1) in a full-length nuclear receptor, which supports a role for AF-1 in SRC-3 recruitment. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6241.map.gz emd_6241.map.gz | 4.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6241-v30.xml emd-6241-v30.xml emd-6241.xml emd-6241.xml | 10.5 KB 10.5 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6241.gif 400_6241.gif 80_6241.gif 80_6241.gif | 40.9 KB 3.9 KB | ||
| Others |  emd_6241_additional_1.map.gz emd_6241_additional_1.map.gz emd_6241_additional_2.map.gz emd_6241_additional_2.map.gz emd_6241_additional_3.map.gz emd_6241_additional_3.map.gz p300-ab1.map.gz p300-ab1.map.gz p300-ab2.map.gz p300-ab2.map.gz | 25.2 MB 1.8 MB 9.8 MB 20 MB 5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6241 http://ftp.pdbj.org/pub/emdb/structures/EMD-6241 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6241 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6241 | HTTPS FTP |
-Related structure data
| Related structure data |  6259C  6260C  6261C  6262C  6263C  6264C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6241.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6241.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ERa/SRC-3/p300 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 6241 additional 1.map
| File | emd_6241_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
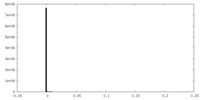
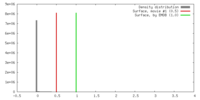
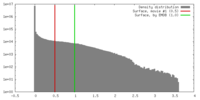
| Density Histograms |
-Supplemental map: emd 6241 additional 2.map
| File | emd_6241_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Supplemental map: emd 6241 additional 3.map
| File | emd_6241_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Supplemental map: p300-ab1.map
| File | p300-ab1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
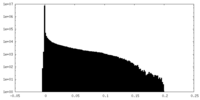
| Density Histograms |
-Supplemental map: p300-ab2.map
| File | p300-ab2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ERa/SRC-3/p300 complex
| Entire | Name: ERa/SRC-3/p300 complex |
|---|---|
| Components |
|
-Supramolecule #1000: ERa/SRC-3/p300 complex
| Supramolecule | Name: ERa/SRC-3/p300 complex / type: sample / ID: 1000 / Number unique components: 3 |
|---|---|
| Molecular weight | Experimental: 800 KDa / Theoretical: 770 KDa / Method: Sedimentation |
-Macromolecule #1: estrogen receptor alpha
| Macromolecule | Name: estrogen receptor alpha / type: protein_or_peptide / ID: 1 / Name.synonym: ERalpha / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Recombinant expression | Organism:  |
-Macromolecule #2: steroid receptor coactivator-3
| Macromolecule | Name: steroid receptor coactivator-3 / type: protein_or_peptide / ID: 2 / Name.synonym: SRC-3 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Recombinant expression | Organism:  |
-Macromolecule #3: p300
| Macromolecule | Name: p300 / type: protein_or_peptide / ID: 3 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: continuous carbon film supported by a 200-mesh R1.2/1.3 Quantifoil grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Method: Blot 1 second prior to plunging. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000 times magnification. |
| Date | Oct 1, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Sampling interval: 15 µm / Number real images: 806 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | Refinements were carried out with a final search angle of 2 degrees using the e2refine.py program. |
|---|---|
| CTF correction | Details: per image |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: OTHER / Software - Name: EMAN2 / Number images used: 18120 |
| Final angle assignment | Details: EMAN2: final search angle 2 degrees |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)