+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of GPCR16-Gi2 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Tas2R16 / GPCR / Gi2 / Salicin / SIGNALING PROTEIN / SIGNALING PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationbitter taste receptor activity / negative regulation of adenylate cyclase-activating adrenergic receptor signaling pathway / negative regulation of calcium ion-dependent exocytosis / detection of chemical stimulus involved in sensory perception of bitter taste / G protein-coupled adenosine receptor signaling pathway / negative regulation of adenylate cyclase activity / ganglioside catabolic process / positive regulation of neural precursor cell proliferation / negative regulation of synaptic transmission / Class C/3 (Metabotropic glutamate/pheromone receptors) ...bitter taste receptor activity / negative regulation of adenylate cyclase-activating adrenergic receptor signaling pathway / negative regulation of calcium ion-dependent exocytosis / detection of chemical stimulus involved in sensory perception of bitter taste / G protein-coupled adenosine receptor signaling pathway / negative regulation of adenylate cyclase activity / ganglioside catabolic process / positive regulation of neural precursor cell proliferation / negative regulation of synaptic transmission / Class C/3 (Metabotropic glutamate/pheromone receptors) / oligosaccharide catabolic process / positive regulation of urine volume / gamma-aminobutyric acid signaling pathway / regulation of calcium ion transport / exo-alpha-sialidase / exo-alpha-sialidase activity / negative regulation of apoptotic signaling pathway / neuronal dense core vesicle / positive regulation of vascular associated smooth muscle cell proliferation / positive regulation of superoxide anion generation / Adenylate cyclase inhibitory pathway / response to nutrient / hippocampal mossy fiber to CA3 synapse / trans-Golgi network / Regulation of insulin secretion / G protein-coupled receptor binding / G protein-coupled receptor activity / G-protein beta/gamma-subunit complex binding / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / G alpha (12/13) signalling events / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / cell body / fibroblast proliferation / Ca2+ pathway / midbody / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction / positive regulation of ERK1 and ERK2 cascade / Extra-nuclear estrogen signaling / cell population proliferation / positive regulation of cell migration / ciliary basal body / G protein-coupled receptor signaling pathway / external side of plasma membrane / cell division / lysosomal membrane / GTPase activity / positive regulation of cell population proliferation / centrosome / synapse / dendrite / GTP binding / protein-containing complex binding / endoplasmic reticulum / signal transduction / extracellular exosome Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.77 Å | |||||||||
 Authors Authors | Wang X / Zhou C / Wu LJ / Hua T / Liu ZJ | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2025 Journal: Cell Rep / Year: 2025Title: Structural basis of β-glucopyranoside salicin recognition by a human bitter taste GPCR. Authors: Xin Wang / Cui Zhou / Weizhen Ao / Lijie Wu / Yiran Wu / Weixiu Xu / Shenhui Liu / Qiwen Tan / Ling Wang / Fei Zhao / Junlin Liu / Yuan Pei / Suwen Zhao / Tian Hua /  Abstract: The human perception of bitterness is mediated by type 2 taste receptors (TAS2Rs), which recognize a broad array of bitter substances with distinct chemical properties. TAS2R16 exhibits a pronounced ...The human perception of bitterness is mediated by type 2 taste receptors (TAS2Rs), which recognize a broad array of bitter substances with distinct chemical properties. TAS2R16 exhibits a pronounced selectivity for β-glucoside-moiety-containing compounds, such as salicin from willow bark. However, the molecular mechanism of moiety-specific recognition and receptor activation in TAS2R16 remains unclear. Here, we present cryoelectron microscopy structures of the salicin-activated human TAS2R16 complexed with gustducin and G and G proteins. The binding mode of salicin with TAS2R16 and the specific interactions of the β-D-glucopyranoside moiety are detailed. Together with molecular docking and mutagenesis data, this study uncovers the structural underpinnings of TAS2R16's group-specific recognition, receptor activation, and subsequent gustducin and G protein coupling. These findings advance our understanding of human bitter taste receptors and provide a foundation for structural modifications of bitter glycosides, opening potential therapeutic applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_62129.map.gz emd_62129.map.gz | 56.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-62129-v30.xml emd-62129-v30.xml emd-62129.xml emd-62129.xml | 24.1 KB 24.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_62129_fsc.xml emd_62129_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_62129.png emd_62129.png | 67.9 KB | ||
| Filedesc metadata |  emd-62129.cif.gz emd-62129.cif.gz | 7.5 KB | ||
| Others |  emd_62129_half_map_1.map.gz emd_62129_half_map_1.map.gz emd_62129_half_map_2.map.gz emd_62129_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-62129 http://ftp.pdbj.org/pub/emdb/structures/EMD-62129 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-62129 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-62129 | HTTPS FTP |
-Related structure data
| Related structure data |  9k6lMC  9kpdC  9kpeC  9kpfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_62129.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_62129.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_62129_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_62129_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : tas2r16 with gi2
| Entire | Name: tas2r16 with gi2 |
|---|---|
| Components |
|
-Supramolecule #1: tas2r16 with gi2
| Supramolecule | Name: tas2r16 with gi2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-2 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.560895 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTVSAEDK AAAERSKMID KNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEDG YSEEECRQYR AVVYSNTIQS IMAIVKAMG NLQIDFADPS RADDARQLFA LSCTAEEQGV LPDDLSGVIR RLWADHGVQA CFGRSREYQL NDSAAYYLND L ERIAQSDY ...String: MGCTVSAEDK AAAERSKMID KNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEDG YSEEECRQYR AVVYSNTIQS IMAIVKAMG NLQIDFADPS RADDARQLFA LSCTAEEQGV LPDDLSGVIR RLWADHGVQA CFGRSREYQL NDSAAYYLND L ERIAQSDY IPTQQDVLRT RVKTTGIVET HFTFKDLHFK MFDVGAQRSE RKKWIHCFEG VTAIIFCVAL SAYDLVLAED EE MNRMHES MKLFDSICNN KWFTDTSIIL FLNKKDLFEE KITHSPLTIC FPEYTGANKY DEAASYIQSK FEDLNKRKDT KEI YTHFTC STDTKNVQFV FDAVTDVIIK NNLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-2 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.728426 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPNG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWNGSSGGG GSGGGGSSGV SGWRLFKKIS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.66682 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAAENLYFQ SHHHHHHHH |
-Macromolecule #5: exo-alpha-sialidase,Taste receptor type 2 member 16,LgBiT
| Macromolecule | Name: exo-alpha-sialidase,Taste receptor type 2 member 16,LgBiT type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO / EC number: exo-alpha-sialidase |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 113.855758 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDAHHHHHH HHHHENLYFQ SAHHHHHHSS GLEVLFQGPP EGAALTEKTD IFESGRNGNP NKDGIKSYR IPALLKTDKG TLIAGADERR LHSSDWGDIG MVIRRSEDNG KTWGDRVTIT NLRDNPKASD PSIGSPVNID M VLVQDPET ...String: MKTIIALSYI FCLVFADYKD DDDAHHHHHH HHHHENLYFQ SAHHHHHHSS GLEVLFQGPP EGAALTEKTD IFESGRNGNP NKDGIKSYR IPALLKTDKG TLIAGADERR LHSSDWGDIG MVIRRSEDNG KTWGDRVTIT NLRDNPKASD PSIGSPVNID M VLVQDPET KRIFSIYDMF PEGKGIFGMS SQKEEAYKKI DGKTYQILYR EGEKGAYTIR ENGTVYTPDG KATDYRVVVD PV KPAYSDK GDLYKGDQLL GNIYFTTNKT SPFRIAKDSY LWMSYSDDDG KTWSAPQDIT PMVKADWMKF LGVGPGTGIV LRN GPHKGR ILIPVYTTNN VSHLDGSQSS RVIYSDDHGK TWHAGEAVND NRQVDGQKIH SSTMNNRRAQ NTESTVVQLN NGDV KLFMR GLTGDLQVAT SKDGGVTWEK DIKRYPQVKD VYVQMSAIHT MHEGKEYIIL SNAGGPKREN GMVHLARVEE NGELT WLKH NPIQKGEFAY NSLQELGNGE YGILYEHTEK GQNAYTLSFR KFNWEFLSKN GSGSGIPIQL TVFFMIIYVL ESLTII VQS SLIVAVLGRE WLQVRRLMPV DMILISLGIS RFCLQWASML NNFCSYFNLN YVLCNLTITW EFFNILTFWL NSLLTVF YC IKVSSFTHHI FLWLRWRILR LFPWILLGCL MITCVTIIPS AIGNYIQIQL LTMEHLPRNS TVTDKLENFH QYQFQAHT V ALVIPFILFL ASTIFLMASL TKQIQHHSTG HCNPSMKARF TALRSLAVLF IVFTSYFLTI LITIIGTLFD KRCWLWVWE AFVYAFILMH STSLMLSSPT LKRILKGKCG SGSGGSGSGG SGSGGSGSGS SGGVFTLEDF VGDWEQTAAY NLDQVLEQGG VSSLLQNLA VSVTPIQRIV RSGENALKID IHVIIPYEGL SADQMAQIEE VFKVVYPVDD HHFKVILPYG TLVIDGVTPN M LNYFGRPY EGIAVFDGKK ITVTGTLWNG NKIIDERLIT PDGSMLFRVT INS UniProtKB: Sialidase A, Taste receptor type 2 member 16 |
-Macromolecule #6: 2-(hydroxymethyl)phenyl beta-D-glucopyranoside
| Macromolecule | Name: 2-(hydroxymethyl)phenyl beta-D-glucopyranoside / type: ligand / ID: 6 / Number of copies: 1 / Formula: SA0 |
|---|---|
| Molecular weight | Theoretical: 286.278 Da |
| Chemical component information | 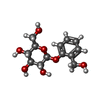 ChemComp-SA0: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOCONTINUUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





































