+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6178 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM map of Mycobacterium tuberculosis 50S ribosome | |||||||||
 Map data Map data | Mycobacterium tuberculosis 50S ribosome | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mycobacterium tuberculosis / ribosome / 50S | |||||||||
| Biological species |  | |||||||||
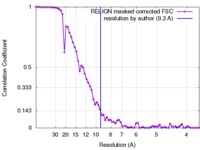
| Method | single particle reconstruction / cryo EM / Resolution: 9.3 Å | |||||||||
 Authors Authors | Yang K / Zhang J | |||||||||
 Citation Citation |  Journal: Structure / Year: 2015 Journal: Structure / Year: 2015Title: Structure of Ribosomal Silencing Factor Bound to Mycobacterium tuberculosis Ribosome. Authors: Xiaojun Li / Qingan Sun / Cai Jiang / Kailu Yang / Li-Wei Hung / Junjie Zhang / James C Sacchettini /  Abstract: The ribosomal silencing factor RsfS slows cell growth by inhibiting protein synthesis during periods of diminished nutrient availability. The crystal structure of Mycobacterium tuberculosis (Mtb) ...The ribosomal silencing factor RsfS slows cell growth by inhibiting protein synthesis during periods of diminished nutrient availability. The crystal structure of Mycobacterium tuberculosis (Mtb) RsfS, together with the cryo-electron microscopy (EM) structure of the large subunit 50S of Mtb ribosome, reveals how inhibition of protein synthesis by RsfS occurs. RsfS binds to the 50S at L14, which, when occupied, blocks the association of the small subunit 30S. Although Mtb RsfS is a dimer in solution, only a single subunit binds to 50S. The overlap between the dimer interface and the L14 binding interface confirms that the RsfS dimer must first dissociate to a monomer in order to bind to L14. RsfS interacts primarily through electrostatic and hydrogen bonding to L14. The EM structure shows extended rRNA density that it is not found in the Escherichia coli ribosome, the most striking of these being the extended RNA helix of H54a. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6178.map.gz emd_6178.map.gz | 8.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6178-v30.xml emd-6178-v30.xml emd-6178.xml emd-6178.xml | 9.1 KB 9.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6178_fsc.xml emd_6178_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  400_6178.gif 400_6178.gif 80_6178.gif 80_6178.gif | 55.7 KB 4.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6178 http://ftp.pdbj.org/pub/emdb/structures/EMD-6178 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6178 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6178 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6178.map.gz / Format: CCP4 / Size: 26.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6178.map.gz / Format: CCP4 / Size: 26.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium tuberculosis 50S ribosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
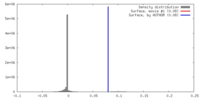
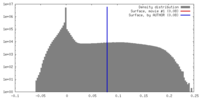
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mycobacterium tuberculosis ribosome 50S
| Entire | Name: Mycobacterium tuberculosis ribosome 50S |
|---|---|
| Components |
|
-Supramolecule #1000: Mycobacterium tuberculosis ribosome 50S
| Supramolecule | Name: Mycobacterium tuberculosis ribosome 50S / type: sample / ID: 1000 / Details: The sample was monodisperse / Oligomeric state: 1 / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 1.6 MDa / Theoretical: 1.6 MDa |
-Supramolecule #1: 50S ribosome
| Supramolecule | Name: 50S ribosome / type: complex / ID: 1 / Recombinant expression: No / Database: NCBI Ribosome-details: ribosome-prokaryote: LSU 50S, LSU RNA 23S, LSU RNA 5S |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.6 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 5 mM HEPES sodium, pH 7.5, 10 mM NH4Cl, 50 mM KCl, 10 mM MgCl2 |
| Grid | Details: 200 mesh R2/2 Quantifoil grid, glow-discharged |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK I |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Average: 100 K |
| Date | Aug 6, 2013 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 165 / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 81081 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 62000 |
| Sample stage | Specimen holder: 626 holder / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)