+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of BTN2A1-BTN3A1-BTN3A3 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | complex / immune / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationButyrophilin (BTN) family interactions / T cell mediated immunity / activated T cell proliferation / regulation of cytokine production / positive regulation of cytokine production / lipid metabolic process / positive regulation of type II interferon production / T cell receptor signaling pathway / adaptive immune response / nuclear body ...Butyrophilin (BTN) family interactions / T cell mediated immunity / activated T cell proliferation / regulation of cytokine production / positive regulation of cytokine production / lipid metabolic process / positive regulation of type II interferon production / T cell receptor signaling pathway / adaptive immune response / nuclear body / signaling receptor binding / external side of plasma membrane / nucleoplasm / membrane / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.07 Å | |||||||||
 Authors Authors | Gao W / Zheng J / Zhu Y / Huang Z | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Immunity / Year: 2025 Journal: Immunity / Year: 2025Title: Phosphoantigen-induced inside-out stabilization of butyrophilin receptor complexes drives dimerization-dependent γδ TCR activation. Authors: Yuwei Zhu / Wenbo Gao / Jianlin Zheng / Ye Bai / Xinyu Tian / Tengjin Huang / Zebin Lu / De Dong / Anqi Zhang / Changyou Guo / Zhiwei Huang /  Abstract: Phosphoantigens (pAgs), produced by infected or cancer cells, trigger the assembly of a membrane receptor complex comprising butyrophilin (BTN) members BTN3A1 and BTN2A1, leading to the activation of ...Phosphoantigens (pAgs), produced by infected or cancer cells, trigger the assembly of a membrane receptor complex comprising butyrophilin (BTN) members BTN3A1 and BTN2A1, leading to the activation of γδ T cells. BTN3A2 or BTN3A3 forms heteromers with BTN3A1, exhibiting higher γδ T cell receptor (TCR)-stimulating activity than BTN3A1 homomers. Cryoelectron microscopy (cryo-EM) structure reveals a pAg-induced BTN2A1-BTN3A1 heterotetramer with a 2:2 stoichiometry, stabilized by interactions between the intracellular B30.2 domains and the extracellular immunoglobulin V (IgV) domains. BTN3A2 or BTN3A3 heterodimerizes with BTN3A1, forming a pAg-induced tetrameric complex with BTN2A1. However, BTN3A1 heterodimers are more stable than BTN3A1 homodimers in this interaction. Cryo-EM reveals that BTN2A1-BTN3A1-BTN3A2 binds two γδ TCR ectodomains, with one being sandwiched between the IgV domains of BTN2A1 and BTN3A2, while the other interacts with the free BTN2A1 IgV in the complex, as evidenced by functional data. Together, our findings uncover the mechanism of ligand-induced inside-out stabilization of BTN receptor complexes for dimeric activation of γδ TCR. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60577.map.gz emd_60577.map.gz | 411.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60577-v30.xml emd-60577-v30.xml emd-60577.xml emd-60577.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_60577.png emd_60577.png | 78.8 KB | ||
| Filedesc metadata |  emd-60577.cif.gz emd-60577.cif.gz | 6.5 KB | ||
| Others |  emd_60577_half_map_1.map.gz emd_60577_half_map_1.map.gz emd_60577_half_map_2.map.gz emd_60577_half_map_2.map.gz | 764.7 MB 764.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60577 http://ftp.pdbj.org/pub/emdb/structures/EMD-60577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60577 | HTTPS FTP |
-Related structure data
| Related structure data |  8zyrMC  8za6C  8za9C  8zaaC  8zabC  8zd4C  9ii6C  9iikC  9j5jC  9j5mC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60577.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60577.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_60577_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_60577_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : complex
| Entire | Name: complex |
|---|---|
| Components |
|
-Supramolecule #1: complex
| Supramolecule | Name: complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Butyrophilin subfamily 2 member A1
| Macromolecule | Name: Butyrophilin subfamily 2 member A1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.826953 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QFIVVGPTDP ILATVGENTT LRCHLSPEKN AEDMEVRWFR SQFSPAVFVY KGGRERTEEQ MEEYRGRTTF VSKDISRGSV ALVIHNITA QENGTYRCYF QEGRSYDEAI LHLVVAGLGS KPLISMRGHE DGGIRLECIS RGWYPKPLTV WRDPYGGVAP A LKEVSMPD ...String: QFIVVGPTDP ILATVGENTT LRCHLSPEKN AEDMEVRWFR SQFSPAVFVY KGGRERTEEQ MEEYRGRTTF VSKDISRGSV ALVIHNITA QENGTYRCYF QEGRSYDEAI LHLVVAGLGS KPLISMRGHE DGGIRLECIS RGWYPKPLTV WRDPYGGVAP A LKEVSMPD ADGLFMVTTA VIIRDKSVRN MSCSINNTLL GQKKESVIFI PESFMPSVSP CAVALPIIVV ILMIPIAVCI YW INKLQKE KKILSGEKEF ERETREIALK ELEKERVQKE EELQVKEKLQ EELRWRRTFL HAVDVVLDPD TAHPDLFLSE DRR SVRRCP FRHLGESVPD NPERFDSQPC VLGRESFASG KHYWEVEVEN VIEWTVGVCR DSVERKGEVL LIPQNGFWTL EMHK GQYRA VSSPDRILPL KESLCRVGVF LDYEAGDVSF YNMRDRSHIY TCPRSAFSVP VRPFFRLGCE DSPIFICPAL TGANG VTVP EEGLTLHRVG THQSL UniProtKB: Butyrophilin subfamily 2 member A1 |
-Macromolecule #2: Butyrophilin subfamily 3 member A1
| Macromolecule | Name: Butyrophilin subfamily 3 member A1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.432809 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QFSVLGPSGP ILAMVGEDAD LPCHLFPTMS AETMELKWVS SSLRQVVNVY ADGKEVEDRQ SAPYRGRTSI LRDGITAGKA ALRIHNVTA SDSGKYLCYF QDGDFYEKAL VELKVAALGS DLHVDVKGYK DGGIHLECRS TGWYPQPQIQ WSNNKGENIP T VEAPVVAD ...String: QFSVLGPSGP ILAMVGEDAD LPCHLFPTMS AETMELKWVS SSLRQVVNVY ADGKEVEDRQ SAPYRGRTSI LRDGITAGKA ALRIHNVTA SDSGKYLCYF QDGDFYEKAL VELKVAALGS DLHVDVKGYK DGGIHLECRS TGWYPQPQIQ WSNNKGENIP T VEAPVVAD GVGLYAVAAS VIMRGSSGEG VSCTIRSSLL GLEKTASISI ADPFFRSAQR WIAALAGTLP VLLLLLGGAG YF LWQQQEE KKTQFRKKKR EQELREMAWS TMKQEQSTRV KLLEELRWRS IQYASRGERH SAYNEWKKAL FKPADVILDP KTA NPILLV SEDQRSVQRA KEPQDLPDNP ERFNWHYCVL GCESFISGRH YWEVEVGDRK EWHIGVCSKN VQRKGWVKMT PENG FWTMG LTDGNKYRTL TEPRTNLKLP KTPKKVGVFL DYETGDISFY NAVDGSHIHT FLDVSFSEAL YPVFRILTLE PTALT ICPA UniProtKB: Butyrophilin subfamily 3 member A1 |
-Macromolecule #3: Butyrophilin subfamily 3 member A3
| Macromolecule | Name: Butyrophilin subfamily 3 member A3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 61.894859 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QFSVLGPSGP ILAMVGEDAD LPCHLFPTMS AETMELRWVS SSLRQVVNVY ADGKEVEDRQ SAPYRGRTSI LRDGITAGKA ALRIHNVTA SDSGKYLCYF QDGDFYEKAL VELKVAALGS DLHIEVKGYE DGGIHLECRS TGWYPQPQIK WSDTKGENIP A VEAPVVAD ...String: QFSVLGPSGP ILAMVGEDAD LPCHLFPTMS AETMELRWVS SSLRQVVNVY ADGKEVEDRQ SAPYRGRTSI LRDGITAGKA ALRIHNVTA SDSGKYLCYF QDGDFYEKAL VELKVAALGS DLHIEVKGYE DGGIHLECRS TGWYPQPQIK WSDTKGENIP A VEAPVVAD GVGLYAVAAS VIMRGSSGGG VSCIIRNSLL GLEKTASISI ADPFFRSAQP WIAALAGTLP ISLLLLAGAS YF LWRQQKE KIALSRETER EREMKEMGYA ATEQEISLRE KLQEELKWRK IQYMARGEKS LAYHEWKMAL FKPADVILDP DTA NAILLV SEDQRSVQRA EEPRDLPDNP ERFEWRYCVL GCENFTSGRH YWEVEVGDRK EWHIGVCSKN VERKKGWVKM TPEN GYWTM GLTDGNKYRA LTEPRTNLKL PEPPRKVGIF LDYETGEISF YNATDGSHIY TFPHASFSEP LYPVFRILTL EPTAL TICP IPKEVESSPD PDLVPDHSLE TPLTPGLANE SGEPQAEVTS LLLPAHPGAE VSPSATTNQN HKLQARTEAL Y UniProtKB: Butyrophilin subfamily 3 member A3 |
-Macromolecule #4: (2E)-4-hydroxy-3-methylbut-2-en-1-yl trihydrogen diphosphate
| Macromolecule | Name: (2E)-4-hydroxy-3-methylbut-2-en-1-yl trihydrogen diphosphate type: ligand / ID: 4 / Number of copies: 1 / Formula: H6P |
|---|---|
| Molecular weight | Theoretical: 262.092 Da |
| Chemical component information | 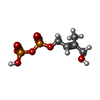 ChemComp-H6P: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































