+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5890 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of MAVS CARD filament | |||||||||
 Map data Map data | Cryo-EM map of MAVS CARD filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Innate immunity / helical filament | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of IP-10 production / regulation of peroxisome organization / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of myeloid dendritic cell cytokine production / CARD domain binding / positive regulation of response to cytokine stimulus / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / protein localization to mitochondrion / positive regulation of type I interferon-mediated signaling pathway / peroxisomal membrane ...positive regulation of IP-10 production / regulation of peroxisome organization / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of myeloid dendritic cell cytokine production / CARD domain binding / positive regulation of response to cytokine stimulus / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / protein localization to mitochondrion / positive regulation of type I interferon-mediated signaling pathway / peroxisomal membrane / TRAF6 mediated IRF7 activation / negative regulation of viral genome replication / type I interferon-mediated signaling pathway / cellular response to exogenous dsRNA / cytoplasmic pattern recognition receptor signaling pathway / TRAF6 mediated NF-kB activation / positive regulation of NLRP3 inflammasome complex assembly / positive regulation of interferon-alpha production / ubiquitin ligase complex / cellular response to interferon-beta / positive regulation of defense response to virus by host / positive regulation of type I interferon production / signaling adaptor activity / activation of innate immune response / positive regulation of interferon-beta production / antiviral innate immune response / positive regulation of interleukin-8 production / protein serine/threonine kinase binding / Negative regulators of DDX58/IFIH1 signaling / protein tetramerization / molecular condensate scaffold activity / positive regulation of protein import into nucleus / DDX58/IFIH1-mediated induction of interferon-alpha/beta / PKR-mediated signaling / positive regulation of interleukin-6 production / Evasion by RSV of host interferon responses / mitochondrial membrane / positive regulation of tumor necrosis factor production / SARS-CoV-1 activates/modulates innate immune responses / Ovarian tumor domain proteases / TRAF3-dependent IRF activation pathway / DNA-binding transcription factor binding / defense response to virus / molecular adaptor activity / protein-macromolecule adaptor activity / mitochondrial outer membrane / positive regulation of canonical NF-kappaB signal transduction / defense response to bacterium / intracellular signal transduction / innate immune response / protein kinase binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses / signal transduction / positive regulation of transcription by RNA polymerase II / mitochondrion / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 9.6 Å | |||||||||
 Authors Authors | Xu H / He X / Zheng H / Huang LJ / Hou F / Yu Z / de la Cruz MJ / Borkowski B / Zhang X / Chen ZJ / Jiang Q-X | |||||||||
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: Structural basis for the prion-like MAVS filaments in antiviral innate immunity. Authors: Hui Xu / Xiaojing He / Hui Zheng / Lily J Huang / Fajian Hou / Zhiheng Yu / Michael Jason de la Cruz / Brian Borkowski / Xuewu Zhang / Zhijian J Chen / Qiu-Xing Jiang /  Abstract: Mitochondrial antiviral signaling (MAVS) protein is required for innate immune responses against RNA viruses. In virus-infected cells MAVS forms prion-like aggregates to activate antiviral signaling ...Mitochondrial antiviral signaling (MAVS) protein is required for innate immune responses against RNA viruses. In virus-infected cells MAVS forms prion-like aggregates to activate antiviral signaling cascades, but the underlying structural mechanism is unknown. Here we report cryo-electron microscopic structures of the helical filaments formed by both the N-terminal caspase activation and recruitment domain (CARD) of MAVS and a truncated MAVS lacking part of the proline-rich region and the C-terminal transmembrane domain. Both structures are left-handed three-stranded helical filaments, revealing specific interfaces between individual CARD subunits that are dictated by electrostatic interactions between neighboring strands and hydrophobic interactions within each strand. Point mutations at multiple locations of these two interfaces impaired filament formation and antiviral signaling. Super-resolution imaging of virus-infected cells revealed rod-shaped MAVS clusters on mitochondria. These results elucidate the structural mechanism of MAVS polymerization, and explain how an α-helical domain uses distinct chemical interactions to form self-perpetuating filaments. DOI: http://dx.doi.org/10.7554/eLife.01489.001. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5890.map.gz emd_5890.map.gz | 297.2 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5890-v30.xml emd-5890-v30.xml emd-5890.xml emd-5890.xml | 12 KB 12 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5890.tif emd_5890.tif | 129.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5890 http://ftp.pdbj.org/pub/emdb/structures/EMD-5890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5890 | HTTPS FTP |
-Related structure data
| Related structure data |  3j6cMC  5891C  4o9fC  4o9lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10014 (Title: MAVS CARD and DeltaProTM filaments / Data size: 35.3 / Data #1: CARD set1 micrographs [micrographs - single frame] / Data #2: CARD set2 micrographs [micrographs - single frame] / Data #3: CARD set3 micrographs [micrographs - single frame] / Data #4: CARD set4 micrographs [micrographs - single frame] EMPIAR-10014 (Title: MAVS CARD and DeltaProTM filaments / Data size: 35.3 / Data #1: CARD set1 micrographs [micrographs - single frame] / Data #2: CARD set2 micrographs [micrographs - single frame] / Data #3: CARD set3 micrographs [micrographs - single frame] / Data #4: CARD set4 micrographs [micrographs - single frame]Data #5: CARD set1 segments [picked particles - multiframe - unprocessed] Data #6: CARD set2 segments [picked particles - multiframe - unprocessed] Data #7: CARD set3 segments [picked particles - multiframe - unprocessed] Data #8: CARD set4 segments [picked particles - multiframe - unprocessed] Data #9: DeltProTM micrographs [micrographs - single frame] Data #10: DeltaProTM segments [picked particles - multiframe - unprocessed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5890.map.gz / Format: CCP4 / Size: 955.1 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5890.map.gz / Format: CCP4 / Size: 955.1 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of MAVS CARD filament | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.333 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
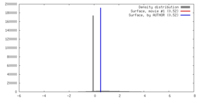
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : MAVS CARD filament
| Entire | Name: MAVS CARD filament |
|---|---|
| Components |
|
-Supramolecule #1000: MAVS CARD filament
| Supramolecule | Name: MAVS CARD filament / type: sample / ID: 1000 / Oligomeric state: polymer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 546 KDa |
-Macromolecule #1: Mitochondrial antiviral-signaling protein (MAVS)
| Macromolecule | Name: Mitochondrial antiviral-signaling protein (MAVS) / type: protein_or_peptide / ID: 1 / Name.synonym: IPS1, KIAA1271, VISA / Oligomeric state: polymer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 13 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK293T / Recombinant plasmid: pcDNA3 Homo sapiens (human) / Recombinant cell: HEK293T / Recombinant plasmid: pcDNA3 |
| Sequence | UniProtKB: Mitochondrial antiviral-signaling protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 20mM Tris-HCl, 50mM NaCl, 1mM DTT |
|---|---|
| Grid | Details: Quantifoil grids coated with thin carbon on top |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FS |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000x magnification |
| Specialist optics | Energy filter - Name: JEOL omega filter / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 35.0 eV |
| Date | Oct 20, 2011 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 61950 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | IHRSR |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 16.8 Å Applied symmetry - Helical parameters - Δ&Phi: 53.6 ° Applied symmetry - Helical parameters - Axial symmetry: C3 (3 fold cyclic) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 9.6 Å / Resolution method: OTHER / Software - Name: Spider, MRC, EMAN, IMAGIC4D Details: Final data were calculated from three separate datasets from three sessions of data collection. The handedness of the map was determined by cryo-electron tomography. |
| CTF correction | Details: Each filament segment |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name: Chimera, SITUS |
| Details | The docking of one X-ray model into a segmented map corresponding to one subunit was first done manually in Chimera, and then optimized using SITUS. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-3j6c: |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)