[English] 日本語
 Yorodumi
Yorodumi- EMDB-5450: Inward-Facing Conformation of the Zinc Transporter YiiP revealed ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5450 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Inward-Facing Conformation of the Zinc Transporter YiiP revealed by Cryo-electron Microscopy | |||||||||
 Map data Map data | This is one unit cell masked from YiiP tubular crystals imaged by cryo-EM | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Zinc transporter / transmembrane protein / electron crystallography / secondary transporter / alternating access mechanism | |||||||||
| Function / homology |  Function and homology information Function and homology informationzinc efflux antiporter activity / cadmium ion transmembrane transporter activity / zinc ion transport / ferrous iron transmembrane transporter activity / intracellular zinc ion homeostasis / membrane => GO:0016020 / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 13.0 Å | |||||||||
 Authors Authors | Coudray N / Valvo S / Hu M / Lasala R / Kim C / Vink M / Zhou M / Provasi D / Filizola M / Tao J ...Coudray N / Valvo S / Hu M / Lasala R / Kim C / Vink M / Zhou M / Provasi D / Filizola M / Tao J / Fang J / Penczek PA / Ubarretxena-Belandia I / Stokes DL | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2013 Journal: Proc Natl Acad Sci U S A / Year: 2013Title: Inward-facing conformation of the zinc transporter YiiP revealed by cryoelectron microscopy. Authors: Nicolas Coudray / Salvatore Valvo / Minghui Hu / Ralph Lasala / Changki Kim / Martin Vink / Ming Zhou / Davide Provasi / Marta Filizola / Juoehi Tao / Jia Fang / Pawel A Penczek / Iban ...Authors: Nicolas Coudray / Salvatore Valvo / Minghui Hu / Ralph Lasala / Changki Kim / Martin Vink / Ming Zhou / Davide Provasi / Marta Filizola / Juoehi Tao / Jia Fang / Pawel A Penczek / Iban Ubarretxena-Belandia / David L Stokes /  Abstract: YiiP is a dimeric Zn(2+)/H(+) antiporter from Escherichia coli belonging to the cation diffusion facilitator family. We used cryoelectron microscopy to determine a 13-Å resolution structure of a ...YiiP is a dimeric Zn(2+)/H(+) antiporter from Escherichia coli belonging to the cation diffusion facilitator family. We used cryoelectron microscopy to determine a 13-Å resolution structure of a YiiP homolog from Shewanella oneidensis within a lipid bilayer in the absence of Zn(2+). Starting from the X-ray structure in the presence of Zn(2+), we used molecular dynamics flexible fitting to build a model consistent with our map. Comparison of the structures suggests a conformational change that involves pivoting of a transmembrane, four-helix bundle (M1, M2, M4, and M5) relative to the M3-M6 helix pair. Although accessibility of transport sites in the X-ray model indicates that it represents an outward-facing state, our model is consistent with an inward-facing state, suggesting that the conformational change is relevant to the alternating access mechanism for transport. Molecular dynamics simulation of YiiP in a lipid environment was used to address the feasibility of this conformational change. Association of the C-terminal domains is the same in both states, and we speculate that this association is responsible for stabilizing the dimer that, in turn, may coordinate the rearrangement of the transmembrane helices. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5450.map.gz emd_5450.map.gz | 74.3 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5450-v30.xml emd-5450-v30.xml emd-5450.xml emd-5450.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5450_1.png emd_5450_1.png | 230 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5450 http://ftp.pdbj.org/pub/emdb/structures/EMD-5450 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5450 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5450 | HTTPS FTP |
-Related structure data
| Related structure data |  3j1zMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5450.map.gz / Format: CCP4 / Size: 19.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5450.map.gz / Format: CCP4 / Size: 19.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is one unit cell masked from YiiP tubular crystals imaged by cryo-EM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.735 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
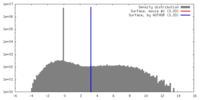
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : YiiP from Shewanella oneidensis in DOPG lipids
| Entire | Name: YiiP from Shewanella oneidensis in DOPG lipids |
|---|---|
| Components |
|
-Supramolecule #1000: YiiP from Shewanella oneidensis in DOPG lipids
| Supramolecule | Name: YiiP from Shewanella oneidensis in DOPG lipids / type: sample / ID: 1000 / Oligomeric state: homodimer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 33 KDa |
-Macromolecule #1: YiiP
| Macromolecule | Name: YiiP / type: protein_or_peptide / ID: 1 / Name.synonym: Ferrous-iron efflux pump (FieF) / Oligomeric state: Dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis MR-1 (bacteria) / Location in cell: Inner membrane Shewanella oneidensis MR-1 (bacteria) / Location in cell: Inner membrane |
| Molecular weight | Theoretical: 33 KDa |
| Recombinant expression | Organism:  |
| Sequence | GO: zinc ion transport, membrane => GO:0016020 / InterPro: Cation efflux protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7 / Details: 20mM TES, 5mM MgCl2, 100mM NaCl, 5mM NaN3 |
| Grid | Details: Holey carbon grids |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 100 K / Instrument: GATAN CRYOPLUNGE 3 / Method: Blot for 2-5 seconds before plunging. |
| Details | Proteins solubilized in DM were mixed with 18:1 dioleoylphosphatidyl glycerol lipids solubilized in DDM at a lipid-to-protein mass ratio of 1. This solution was dialyzed at room temperature for two weeks. Crystal cell parameters were a=57.5, b=34.0, c=100.0, alpha=90, beta=90, gamma=85.3. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: Objective astigmatism corrected at 200,000 times magnification |
| Date | Feb 10, 2009 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 14 µm / Number real images: 19 / Average electron dose: 10 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 51190 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.1 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.6 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Indexing of the helical symmetry was done by comparing the positions of individual layer lines in Fourier space with the outer radius of the tube in real space. IHRSR method used to calculate the 3D structure. |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 17.1 Å Applied symmetry - Helical parameters - Δ&Phi: 56.4 ° Applied symmetry - Helical parameters - Axial symmetry: C3 (3 fold cyclic) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 13.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPARX, EMIP |
| CTF correction | Details: CTF to the whole micrograph |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - #0 - Chain ID: A / Chain - #1 - Chain ID: C |
|---|---|
| Details | Protocol: MDFF. An initial homology model of YiiP from S. oneidensis was built with MODELLER 9v7 using the X-ray crystal structure of YiiP from E. coli (PDB entry 3H90) as a template. This model included 9 residues at the N-terminus and 4 residues at the C-terminus that were not present in the X-ray structure. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-3j1z: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)