[English] 日本語
 Yorodumi
Yorodumi- EMDB-5307: Three-dimensional cryo-electron microscopy density map of the 70S... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5307 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Three-dimensional cryo-electron microscopy density map of the 70S ribosome from Mycobacterium smegmatis | |||||||||
 Map data Map data | Reconstruction of 70S ribosome from Mycobacterium smegmatis | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cryoEM / prokaryotic / ribosome / 3D-structure | |||||||||
| Biological species |  Mycobacterium smegmatis (bacteria) Mycobacterium smegmatis (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 12.0 Å | |||||||||
 Authors Authors | Shasmal M / Sengupta J | |||||||||
 Citation Citation |  Journal: PLoS One / Year: 2012 Journal: PLoS One / Year: 2012Title: Structural diversity in bacterial ribosomes: mycobacterial 70S ribosome structure reveals novel features. Authors: Manidip Shasmal / Jayati Sengupta /  Abstract: Here we present analysis of a 3D cryo-EM map of the 70S ribosome from Mycobacterium smegmatis, a saprophytic cousin of the etiological agent of tuberculosis in humans, Mycobacterium tuberculosis. In ...Here we present analysis of a 3D cryo-EM map of the 70S ribosome from Mycobacterium smegmatis, a saprophytic cousin of the etiological agent of tuberculosis in humans, Mycobacterium tuberculosis. In comparison with the 3D structures of other prokaryotic ribosomes, the density map of the M. smegmatis 70S ribosome reveals unique structural features and their relative orientations in the ribosome. Dramatic changes in the periphery due to additional rRNA segments and extra domains of some of the peripheral ribosomal proteins like S3, S5, S16, L17, L25, are evident. One of the most notable features appears in the large subunit near L1 stalk as a long helical structure next to helix 54 of the 23S rRNA. The sharp upper end of this structure is located in the vicinity of the mRNA exit channel. Although the M. smegmatis 70S ribosome possesses conserved core structure of bacterial ribosome, the new structural features, unveiled in this study, demonstrates diversity in the 3D architecture of bacterial ribosomes. We postulate that the prominent helical structure related to the 23S rRNA actively participates in the mechanisms of translation in mycobacteria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5307.map.gz emd_5307.map.gz | 34.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5307-v30.xml emd-5307-v30.xml emd-5307.xml emd-5307.xml | 9.3 KB 9.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5307_1.tif emd_5307_1.tif | 207.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5307 http://ftp.pdbj.org/pub/emdb/structures/EMD-5307 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5307 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5307 | HTTPS FTP |
-Validation report
| Summary document |  emd_5307_validation.pdf.gz emd_5307_validation.pdf.gz | 78 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5307_full_validation.pdf.gz emd_5307_full_validation.pdf.gz | 77.1 KB | Display | |
| Data in XML |  emd_5307_validation.xml.gz emd_5307_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5307 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5307 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5307 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5307 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5307.map.gz / Format: CCP4 / Size: 37 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5307.map.gz / Format: CCP4 / Size: 37 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of 70S ribosome from Mycobacterium smegmatis | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.69 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
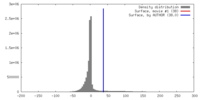
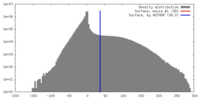
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 70S ribosome from Mycobacterium smegmatis
| Entire | Name: 70S ribosome from Mycobacterium smegmatis |
|---|---|
| Components |
|
-Supramolecule #1000: 70S ribosome from Mycobacterium smegmatis
| Supramolecule | Name: 70S ribosome from Mycobacterium smegmatis / type: sample / ID: 1000 / Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 2.5 MDa |
-Supramolecule #1: 70S ribosome with P-site tRNA
| Supramolecule | Name: 70S ribosome with P-site tRNA / type: complex / ID: 1 / Name.synonym: 70S ribosome / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:  Mycobacterium smegmatis (bacteria) Mycobacterium smegmatis (bacteria) |
| Molecular weight | Theoretical: 2.5 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 20 mM Tris-HCl, pH 7.5, 10 mM magnesium acetate, 30 mM ammonium chloride, 5 mM 2-mercaptoethanol |
|---|---|
| Grid | Details: Holey grid |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 94 K / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: Cryo plunger. Rapid-freezing in liquid ethane |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Nov 2, 2010 |
| Image recording | Category: CCD / Film or detector model: GENERIC CCD / Digitization - Sampling interval: 15 µm / Number real images: 269 / Average electron dose: 15 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 88466 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 89000 |
| Sample stage | Specimen holder: Cryo transfer / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: CTF correction of 3D map |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 12.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPIDER / Number images used: 1 |
-Atomic model buiding 1
| Initial model | PDB ID:  2i2u |
|---|---|
| Software | Name: PyMol |
| Details | Manual fitting |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
-Atomic model buiding 2
| Initial model | PDB ID:  2i2v |
|---|---|
| Software | Name: PyMol |
| Details | Manual fitting |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)