[English] 日本語
 Yorodumi
Yorodumi- EMDB-43085: Symmetry expanded map of 2 gamma-tubulins bound to 2 alpha tubuli... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Symmetry expanded map of 2 gamma-tubulins bound to 2 alpha tubulins in gamma tubulin ring complex capped microtubule end. | |||||||||
 Map data Map data | Symmetry expanded map of gamma-tubulin:gamma tubulin laterally in contact with each other together with the proximal alpha-tubulins in gamma tubulin ring complex upon microtubule nucleation. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | gamma tubulin ring complex / CELL CYCLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationPost-chaperonin tubulin folding pathway / Cilium Assembly / cytoskeleton-dependent intracellular transport / Carboxyterminal post-translational modifications of tubulin / Microtubule-dependent trafficking of connexons from Golgi to the plasma membrane / Sealing of the nuclear envelope (NE) by ESCRT-III / Intraflagellar transport / Formation of tubulin folding intermediates by CCT/TriC / Gap junction assembly / Kinesins ...Post-chaperonin tubulin folding pathway / Cilium Assembly / cytoskeleton-dependent intracellular transport / Carboxyterminal post-translational modifications of tubulin / Microtubule-dependent trafficking of connexons from Golgi to the plasma membrane / Sealing of the nuclear envelope (NE) by ESCRT-III / Intraflagellar transport / Formation of tubulin folding intermediates by CCT/TriC / Gap junction assembly / Kinesins / Assembly and cell surface presentation of NMDA receptors / COPI-independent Golgi-to-ER retrograde traffic / COPI-dependent Golgi-to-ER retrograde traffic / Recycling pathway of L1 / RHOH GTPase cycle / microtubule-based process / RHO GTPases activate IQGAPs / Hedgehog 'off' state / COPI-mediated anterograde transport / cytoplasmic microtubule / Activation of AMPK downstream of NMDARs / MHC class II antigen presentation / Recruitment of NuMA to mitotic centrosomes / Mitotic Prometaphase / cellular response to interleukin-4 / EML4 and NUDC in mitotic spindle formation / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / Resolution of Sister Chromatid Cohesion / Translocation of SLC2A4 (GLUT4) to the plasma membrane / RHO GTPases Activate Formins / PKR-mediated signaling / structural constituent of cytoskeleton / microtubule cytoskeleton organization / HCMV Early Events / Aggrephagy / The role of GTSE1 in G2/M progression after G2 checkpoint / Separation of Sister Chromatids / mitotic cell cycle / double-stranded RNA binding / microtubule cytoskeleton / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / microtubule / cilium / cell division / GTPase activity / ubiquitin protein ligase binding / GTP binding / structural molecule activity / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Trichoplusia ni (cabbage looper) / Trichoplusia ni (cabbage looper) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
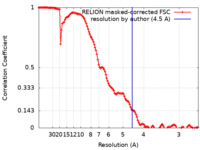
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Aher A / Urnavicius L / Kapoor TM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structure of the γ-tubulin ring complex-capped microtubule. Authors: Amol Aher / Linas Urnavicius / Allen Xue / Kasahun Neselu / Tarun M Kapoor /  Abstract: Microtubules are composed of α-tubulin and β-tubulin dimers positioned head-to-tail to form protofilaments that associate laterally in varying numbers. It is not known how cellular microtubules ...Microtubules are composed of α-tubulin and β-tubulin dimers positioned head-to-tail to form protofilaments that associate laterally in varying numbers. It is not known how cellular microtubules assemble with the canonical 13-protofilament architecture, resulting in micrometer-scale α/β-tubulin tracks for intracellular transport that align with, rather than spiral along, the long axis of the filament. We report that the human ~2.3 MDa γ-tubulin ring complex (γ-TuRC), an essential regulator of microtubule formation that contains 14 γ-tubulins, selectively nucleates 13-protofilament microtubules. Cryogenic electron microscopy reconstructions of γ-TuRC-capped microtubule minus ends reveal the extensive intra-domain and inter-domain motions of γ-TuRC subunits that accommodate luminal bridge components and establish lateral and longitudinal interactions between γ-tubulins and α-tubulins. Our structures suggest that γ-TuRC, an inefficient nucleation template owing to its splayed conformation, can transform into a compacted cap at the microtubule minus end and set the lattice architecture of cellular microtubules. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43085.map.gz emd_43085.map.gz | 6.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43085-v30.xml emd-43085-v30.xml emd-43085.xml emd-43085.xml | 18.9 KB 18.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43085_fsc.xml emd_43085_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_43085.png emd_43085.png | 51.5 KB | ||
| Filedesc metadata |  emd-43085.cif.gz emd-43085.cif.gz | 6.7 KB | ||
| Others |  emd_43085_half_map_1.map.gz emd_43085_half_map_1.map.gz emd_43085_half_map_2.map.gz emd_43085_half_map_2.map.gz | 140.9 MB 140.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43085 http://ftp.pdbj.org/pub/emdb/structures/EMD-43085 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43085 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43085 | HTTPS FTP |
-Related structure data
| Related structure data |  8va2MC  8vrdC  8vrjC  8vrkC  8vt7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43085.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43085.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Symmetry expanded map of gamma-tubulin:gamma tubulin laterally in contact with each other together with the proximal alpha-tubulins in gamma tubulin ring complex upon microtubule nucleation. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Symmetry expanded map of gamma-tubulin:gamma tubulin laterally in...
| File | emd_43085_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Symmetry expanded map of gamma-tubulin:gamma tubulin laterally in contact with each other together with the proximal alpha-tubulins in gamma tubulin ring complex upon microtubule nucleation. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Symmetry expanded map of gamma-tubulin:gamma tubulin laterally in...
| File | emd_43085_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Symmetry expanded map of gamma-tubulin:gamma tubulin laterally in contact with each other together with the proximal alpha-tubulins in gamma tubulin ring complex upon microtubule nucleation. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : gamma-tubulin and alpha-tubulin
| Entire | Name: gamma-tubulin and alpha-tubulin |
|---|---|
| Components |
|
-Supramolecule #1: gamma-tubulin and alpha-tubulin
| Supramolecule | Name: gamma-tubulin and alpha-tubulin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
-Macromolecule #1: Tubulin alpha-1B chain
| Macromolecule | Name: Tubulin alpha-1B chain / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.019297 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIHHHHHHGG GDDSFNTFFS ETGAGKHVPR AVFVDLEPTV IDEVRTGTY RQLFHPEQLI TGKEDAANNY ARGHYTIGKE IIDLVLDRIR KLADQCTGLQ GFLVFHSFGG GTGSGFTSLL M ERLSVDYG ...String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIHHHHHHGG GDDSFNTFFS ETGAGKHVPR AVFVDLEPTV IDEVRTGTY RQLFHPEQLI TGKEDAANNY ARGHYTIGKE IIDLVLDRIR KLADQCTGLQ GFLVFHSFGG GTGSGFTSLL M ERLSVDYG KKSKLEFSIY PAPQVSTAVV EPYNSILTTH TTLEHSDCAF MVDNEAIYDI CRRNLDIERP TYTNLNRLIS QI VSSITAS LRFDGALNVD LTDFQTNLVP YPRIHFPLAT YAPVISAEKA YHEQLSVAEI TNACFEPANQ MVKCDPRHGK YMA CCLLYR GDVVPKDVNA AIATIKTKRS IQFVDWCPTG FKVGINYQPP TVVPGGDLAK VQRAVCMLSN TTAIAEAWAR LDHK FDLMY AKRAFVHWYV GEGMEEGEFS EAREDMAALE KDYEEVGVDS VEGEGEEEGE EY UniProtKB: Tubulin alpha-1B chain |
-Macromolecule #2: Tubulin gamma-1 chain
| Macromolecule | Name: Tubulin gamma-1 chain / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.019539 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MPREIITLQL GQCGNQIGFE FWKQLCAEHG ISPEGIVEEF ATEGTDRKDV FFYQADDEHY IPRAVLLDLE PRVIHSILNS PYAKLYNPE NIYLSEHGGG AGNNWASGFS QGEKIHEDIF DIIDREADGS DSLEGFVLCH SIAGGTGSGL GSYLLERLND R YPKKLVQT ...String: MPREIITLQL GQCGNQIGFE FWKQLCAEHG ISPEGIVEEF ATEGTDRKDV FFYQADDEHY IPRAVLLDLE PRVIHSILNS PYAKLYNPE NIYLSEHGGG AGNNWASGFS QGEKIHEDIF DIIDREADGS DSLEGFVLCH SIAGGTGSGL GSYLLERLND R YPKKLVQT YSVFPNQDEM SDVVVQPYNS LLTLKRLTQN ADCVVVLDNT ALNRIATDRL HIQNPSFSQI NQLVSTIMSA ST TTLRYPG YMNNDLIGLI ASLIPTPRLH FLMTGYTPLT TDQSVASVRK TTVLDVMRRL LQPKNVMVST GRDRQTNHCY IAI LNIIQG EVDPTQVHKS LQRIRERKLA NFIPWGPASI QVALSRKSPY LPSAHRVSGL MMANHTSISS LFERTCRQYD KLRK REAFL EQFRKEDMFK DNFDEMDTSR EIVQQLIDEY HAATRP |
-Macromolecule #3: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 2 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 2 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)