+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of LRRK2 bound to type I inhibitor GNE-7915 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryo-EM / Parkinson's disease / Kinase / LRRK2 / type I inhibitor / HYDROLASE / HYDROLASE-HYDROLASE INHIBITOR complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of multicellular organismal process / intracellular organelle lumen / Wnt signalosome assembly / GTP-dependent protein kinase activity / protein localization to endoplasmic reticulum exit site / animal organ development / regulation of intracellular signal transduction / regulation of dopamine receptor signaling pathway / regulation of vesicle-mediated transport / regulation of dendritic spine morphogenesis ...regulation of multicellular organismal process / intracellular organelle lumen / Wnt signalosome assembly / GTP-dependent protein kinase activity / protein localization to endoplasmic reticulum exit site / animal organ development / regulation of intracellular signal transduction / regulation of dopamine receptor signaling pathway / regulation of vesicle-mediated transport / regulation of dendritic spine morphogenesis / endoplasmic reticulum organization / Wnt signalosome / regulation of canonical Wnt signaling pathway / Golgi organization / protein kinase A binding / exploration behavior / positive regulation of protein metabolic process / regulation of proteolysis / endoplasmic reticulum exit site / anatomical structure morphogenesis / negative regulation of signal transduction / phagocytic vesicle / vesicle-mediated transport / positive regulation of autophagy / GTPase activator activity / regulation of membrane potential / modulation of chemical synaptic transmission / autophagy / small GTPase binding / neuron differentiation / terminal bouton / protein autophosphorylation / synaptic vesicle membrane / signaling receptor complex adaptor activity / regulation of gene expression / cytoskeleton / protein phosphorylation / perikaryon / mitochondrial outer membrane / non-specific serine/threonine protein kinase / cell surface receptor signaling pathway / lysosome / endosome / intracellular signal transduction / membrane raft / Golgi membrane / hydrolase activity / dendrite / endoplasmic reticulum membrane / GTP binding / ATP binding / identical protein binding / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Zhu H / Sun J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2024 Journal: Cell Discov / Year: 2024Title: Pharmacology of LRRK2 with type I and II kinase inhibitors revealed by cryo-EM. Authors: Hanwen Zhu / Patricia Hixson / Wen Ma / Ji Sun /  Abstract: LRRK2 is one of the most promising drug targets for Parkinson's disease. Though type I kinase inhibitors of LRRK2 are under clinical trials, alternative strategies like type II inhibitors are being ...LRRK2 is one of the most promising drug targets for Parkinson's disease. Though type I kinase inhibitors of LRRK2 are under clinical trials, alternative strategies like type II inhibitors are being actively pursued due to the potential undesired effects of type I inhibitors. Currently, a robust method for LRRK2-inhibitor structure determination to guide structure-based drug discovery is lacking, and inhibition mechanisms of available compounds are also unclear. Here we present near-atomic-resolution structures of LRRK2 with type I (LRRK2-IN-1 and GNE-7915) and type II (rebastinib, ponatinib, and GZD-824) inhibitors, uncovering the structural basis of LRRK2 inhibition and conformational plasticity of the kinase domain with molecular dynamics (MD) simulations. Type I and II inhibitors bind to LRRK2 in active-like and inactive conformations, so LRRK2-inhibitor complexes further reveal general structural features associated with LRRK2 activation. Our study provides atomic details of LRRK2-inhibitor interactions and a framework for understanding LRRK2 activation and for rational drug design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41982.map.gz emd_41982.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41982-v30.xml emd-41982-v30.xml emd-41982.xml emd-41982.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41982.png emd_41982.png | 33.8 KB | ||
| Filedesc metadata |  emd-41982.cif.gz emd-41982.cif.gz | 6.1 KB | ||
| Others |  emd_41982_half_map_1.map.gz emd_41982_half_map_1.map.gz emd_41982_half_map_2.map.gz emd_41982_half_map_2.map.gz | 59.5 MB 59.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41982 http://ftp.pdbj.org/pub/emdb/structures/EMD-41982 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41982 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41982 | HTTPS FTP |
-Related structure data
| Related structure data |  8u7hMC  8fo7C  8u7lC  8u8aC  8u8bC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41982.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41982.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.4895 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41982_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_41982_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : LRRK2-GNE-7915
| Entire | Name: LRRK2-GNE-7915 |
|---|---|
| Components |
|
-Supramolecule #1: LRRK2-GNE-7915
| Supramolecule | Name: LRRK2-GNE-7915 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 137 kDa/nm |
-Macromolecule #1: non-specific serine/threonine protein kinase
| Macromolecule | Name: non-specific serine/threonine protein kinase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 136.843469 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: KKAVPYNRMK LMIVGNTGSG KTTLLQQLMK TKKSDLGMQS ATVGIDVKDW PIQIRDKRKR DLVLNVWDFA GREEFYSTHP HFMTQRALY LAVYDLSKGQ AEVDAMKPWL FNIKARASSS PVILVGTHLD VSDEKQRKAC MSKITKELLN KRGFPAIRDY H FVNATEES ...String: KKAVPYNRMK LMIVGNTGSG KTTLLQQLMK TKKSDLGMQS ATVGIDVKDW PIQIRDKRKR DLVLNVWDFA GREEFYSTHP HFMTQRALY LAVYDLSKGQ AEVDAMKPWL FNIKARASSS PVILVGTHLD VSDEKQRKAC MSKITKELLN KRGFPAIRDY H FVNATEES DALAKLRKTI INESLNFKIR DQLVVGQLIP DCYVELEKII LSERKNVPIE FPVIDRKRLL QLVRENQLQL DE NELPHAV HFLNESGVLL HFQDPALQLS DLYFVEPKWL CKIMAQILTV KVEGCPKHPK GIISRRDVEK FLSKKRKFPK NYM TQYFKL LEKFQIALPI GEEYLLVPSS LSDHRPVIEL PHCENSEIII RLYEMPYFPM GFWSRLINRL LEISPYMLSG RERA LRPNR RYWRQGIYLN WSPEAYCLVG SEVLDNHPES FLKITVPSCR KGCILLGQVV DHIDSLMEEW FPGLLEIDIC GEGET LLKK WALYSFNDGE EHQKILLDDL MKKAEEGDLL VNPDQPRLTI PISQIAPDLI LADLPRNIML NNDELEFEQA PEFLLG DGS FGSVYRAAYE GEEVAVKIFN KHTSLRLLRQ ELVVLCHLHH PSLISLLAAG IRPRMLVMEL ASKGSLDRLL QQDKASL TR TLQHRIALHV ADGLRYLHSA MIIYRDLKPH NVLLFTLYPN AAIIAKIADY GIAQYCCRMG IKTSEGTPGF RAPEVARG N VIYNQQADVY SFGLLLYDIL TTGGRIVEGL KFPNEFDELE IQGKLPDPVK EYGCAPWPMV EKLIKQCLKE NPQERPTSA QVFDILNSAE LVCLTRRILL PKNVIVECMV ATHHNSRNAS IWLGCGHTDR GQLSFLDLNT EGYTSEEVAD SRILCLALVH LPVEKESWI VSGTQSGTLL VINTEDGKKR HTLEKMTDSV TCLYCNSFSK QSKQKNFLLV GTADGKLAIF EDKTVKLKGA A PLKILNIG NVSTPLMCLS ESTNSTERNV MWGGCGTKIF SFSNDFTIQK LIETRTSQLF SYAAFSDSNI ITVVVDTALY IA KQNSPVV EVWDKKTEKL CGLIDCVHFL REVTVKENKE SKHKTSYSGR VKTLCLQKNT ALWIGTGGGH ILLLDLSTRR LIR VIYNFC NSVRVMMTAQ LGSLKNVMLV LGYNRKNTEG TQKQKEIQSC LTVWDINLPH EVQNLEKHIE VRKELAEKMR RTSV E UniProtKB: Leucine-rich repeat serine/threonine-protein kinase 2 |
-Macromolecule #2: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 1 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Macromolecule #3: [4-[[4-(ethylamino)-5-(trifluoromethyl)pyrimidin-2-yl]amino]-2-fl...
| Macromolecule | Name: [4-[[4-(ethylamino)-5-(trifluoromethyl)pyrimidin-2-yl]amino]-2-fluoranyl-5-methoxy-phenyl]-morpholin-4-yl-methanone type: ligand / ID: 3 / Number of copies: 1 / Formula: A0T |
|---|---|
| Molecular weight | Theoretical: 443.395 Da |
| Chemical component information | 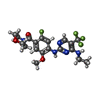 ChemComp-A0T: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 62.08 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 95644 |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)