[English] 日本語
 Yorodumi
Yorodumi- EMDB-38451: Structure of native tri-heteromeric GluN1-GluN2A-GluN2B NMDA rece... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of native tri-heteromeric GluN1-GluN2A-GluN2B NMDA receptor in rat cortex and hippocampus | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MEMBRANE PROTEIN / native NMDA receptor / adult rat cartex & hippocampus / GluN2A / GluN2B / MEMBRANE PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of response to alcohol / response to ammonium ion / neurotransmitter receptor transport, plasma membrane to endosome / cellular response to corticosterone stimulus / cellular response to magnesium starvation / receptor recycling / sensory organ development / response to environmental enrichment / cellular response to curcumin / directional locomotion ...regulation of response to alcohol / response to ammonium ion / neurotransmitter receptor transport, plasma membrane to endosome / cellular response to corticosterone stimulus / cellular response to magnesium starvation / receptor recycling / sensory organ development / response to environmental enrichment / cellular response to curcumin / directional locomotion / regulation of cAMP/PKA signal transduction / pons maturation / EPHB-mediated forward signaling / Assembly and cell surface presentation of NMDA receptors / auditory behavior / positive regulation of Schwann cell migration / regulation of cell communication / sensitization / cellular response to magnesium ion / olfactory learning / response to other organism / response to hydrogen sulfide / serotonin metabolic process / dendritic branch / fear response / conditioned taste aversion / response to methylmercury / protein localization to postsynaptic membrane / regulation of ARF protein signal transduction / regulation of respiratory gaseous exchange / apical dendrite / response to manganese ion / transmitter-gated monoatomic ion channel activity / suckling behavior / sleep / interleukin-1 receptor binding / positive regulation of inhibitory postsynaptic potential / regulation of NMDA receptor activity / response to carbohydrate / cellular response to lipid / propylene metabolic process / response to glycine / dendritic spine organization / locomotion / cellular response to dsRNA / RAF/MAP kinase cascade / negative regulation of dendritic spine maintenance / response to amine / response to growth hormone / heterocyclic compound binding / neurotransmitter receptor complex / Synaptic adhesion-like molecules / response to glycoside / regulation of monoatomic cation transmembrane transport / NMDA glutamate receptor activity / voltage-gated monoatomic cation channel activity / NMDA selective glutamate receptor complex / glutamate binding / glutamate receptor signaling pathway / ligand-gated sodium channel activity / positive regulation of glutamate secretion / neuromuscular process / regulation of axonogenesis / calcium ion transmembrane import into cytosol / regulation of dendrite morphogenesis / male mating behavior / regulation of synapse assembly / protein heterotetramerization / response to morphine / small molecule binding / spinal cord development / glycine binding / receptor clustering / startle response / dopamine metabolic process / cellular response to zinc ion / positive regulation of reactive oxygen species biosynthetic process / response to lithium ion / parallel fiber to Purkinje cell synapse / regulation of MAPK cascade / monoatomic ion channel complex / behavioral response to pain / regulation of postsynaptic membrane potential / monoatomic cation transmembrane transport / action potential / response to electrical stimulus / positive regulation of calcium ion transport into cytosol / extracellularly glutamate-gated ion channel activity / modulation of excitatory postsynaptic potential / cellular response to glycine / associative learning / positive regulation of dendritic spine maintenance / response to light stimulus / response to magnesium ion / positive regulation of protein targeting to membrane / Unblocking of NMDA receptors, glutamate binding and activation / regulation of neuronal synaptic plasticity / monoatomic cation transport / glutamate receptor binding / conditioned place preference Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Zhang M / Feng J / Li Y / Zhu S | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2025 Journal: Cell / Year: 2025Title: Assembly and architecture of endogenous NMDA receptors in adult cerebral cortex and hippocampus. Authors: Ming Zhang / Juan Feng / Chun Xie / Nan Song / Chaozhi Jin / Jian Wang / Qun Zhao / Lihua Zhang / Boshuang Wang / Yidi Sun / Fei Guo / Yang Li / Shujia Zhu /  Abstract: The cerebral cortex and hippocampus are crucial brain regions for learning and memory, which depend on activity-induced synaptic plasticity involving N-methyl-ᴅ-aspartate receptors (NMDARs). ...The cerebral cortex and hippocampus are crucial brain regions for learning and memory, which depend on activity-induced synaptic plasticity involving N-methyl-ᴅ-aspartate receptors (NMDARs). However, subunit assembly and molecular architecture of endogenous NMDARs (eNMDARs) in the brain remain elusive. Using conformation- and subunit-dependent antibodies, we purified eNMDARs from adult rat cerebral cortex and hippocampus. Three major subtypes of GluN1-N2A-N2B, GluN1-N2B, and GluN1-N2A eNMDARs were resolved by cryoelectron microscopy (cryo-EM) at the resolution up to 4.2 Å. The particle ratio of these three subtypes was 9:7:4, indicating that about half of GluN2A and GluN2B subunits are incorporated into the tri-heterotetramers. Structural analysis revealed the asymmetric architecture of the GluN1-N2A-N2B receptor throughout the extracellular to the transmembrane layers. Moreover, the conformational variations between GluN1-N2B and GluN1-N2A-N2B receptors revealed the distinct biophysical properties across different eNMDAR subtypes. Our findings imply the structural and functional complexity of eNMDARs and shed light on structure-based therapeutic design targeting these eNMDARs in vivo. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38451.map.gz emd_38451.map.gz | 111.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38451-v30.xml emd-38451-v30.xml emd-38451.xml emd-38451.xml | 32.7 KB 32.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38451_fsc.xml emd_38451_fsc.xml | 14.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_38451.png emd_38451.png | 141.2 KB | ||
| Masks |  emd_38451_msk_1.map emd_38451_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-38451.cif.gz emd-38451.cif.gz | 9.7 KB | ||
| Others |  emd_38451_half_map_1.map.gz emd_38451_half_map_1.map.gz emd_38451_half_map_2.map.gz emd_38451_half_map_2.map.gz | 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38451 http://ftp.pdbj.org/pub/emdb/structures/EMD-38451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38451 | HTTPS FTP |
-Related structure data
| Related structure data |  8xlkMC  8xljC  8xllC  9jnnC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38451.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38451.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.071 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_38451_msk_1.map emd_38451_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_38451_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_38451_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Native tri-heteromeric GluN1-GluN2A-GluN2B NMDA receptor in rat c...
+Supramolecule #1: Native tri-heteromeric GluN1-GluN2A-GluN2B NMDA receptor in rat c...
+Supramolecule #2: NMDA receptor
+Supramolecule #3: Fab 4F11
+Supramolecule #4: Fab 28C
+Supramolecule #5: Fab2
+Supramolecule #6: Fab
+Macromolecule #1: Glutamate receptor ionotropic, NMDA 1
+Macromolecule #2: Glutamate receptor ionotropic, NMDA 2A
+Macromolecule #3: Glutamate receptor ionotropic, NMDA 2B
+Macromolecule #4: Heavy Chain of GluN1 Fab, 4F11
+Macromolecule #5: Light Chain of GluN1 Fab, 4F11
+Macromolecule #6: Heavy Chain of GluN2A Fab, 28C
+Macromolecule #7: Light Chain of GluN2A Fab, 28C
+Macromolecule #8: Heavy Chain of GluN2B Fab2
+Macromolecule #9: Light Chain of GluN2B Fab2
+Macromolecule #12: 2-acetamido-2-deoxy-beta-D-glucopyranose
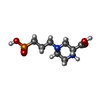
+Macromolecule #13: (2R)-4-(3-phosphonopropyl)piperazine-2-carboxylic acid
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | tissue |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-10 (5k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)












































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)