+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Gq bound FZD1 in ligand-free state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FZD1-Gq / class-F / Frizzled receptor / Complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmuscular septum morphogenesis / regulation of mesenchymal stem cell differentiation / autocrine signaling / astrocyte-dopaminergic neuron signaling / Fatty Acids bound to GPR40 (FFAR1) regulate insulin secretion / presynapse assembly / Acetylcholine regulates insulin secretion / Wnt receptor activity / sensory perception of itch / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway ...muscular septum morphogenesis / regulation of mesenchymal stem cell differentiation / autocrine signaling / astrocyte-dopaminergic neuron signaling / Fatty Acids bound to GPR40 (FFAR1) regulate insulin secretion / presynapse assembly / Acetylcholine regulates insulin secretion / Wnt receptor activity / sensory perception of itch / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / membranous septum morphogenesis / non-canonical Wnt signaling pathway / endothelial cell differentiation / Wnt-protein binding / PLC beta mediated events / phospholipase C-activating serotonin receptor signaling pathway / regulation of platelet activation / entrainment of circadian clock / midbrain dopaminergic neuron differentiation / hard palate development / frizzled binding / negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway / PCP/CE pathway / Class B/2 (Secretin family receptors) / Wnt signalosome / Disassembly of the destruction complex and recruitment of AXIN to the membrane / regulation of canonical Wnt signaling pathway / glutamate receptor signaling pathway / mast cell degranulation / outflow tract morphogenesis / adenylate cyclase-activating G protein-coupled bile acid receptor signaling pathway / adenylate cyclase-activating serotonin receptor signaling pathway / regulation of skeletal muscle contraction / phototransduction, visible light / PKA activation in glucagon signalling / negative regulation of BMP signaling pathway / hair follicle placode formation / developmental growth / intracellular transport / positive regulation of osteoblast differentiation / photoreceptor outer segment / canonical Wnt signaling pathway / regulation of presynapse assembly / D1 dopamine receptor binding / postsynaptic cytosol / vascular endothelial cell response to laminar fluid shear stress / renal water homeostasis / activation of adenylate cyclase activity / Hedgehog 'off' state / cellular response to acidic pH / adenylate cyclase-activating adrenergic receptor signaling pathway / hormone-mediated signaling pathway / cellular response to glucagon stimulus / intracellular glucose homeostasis / Turbulent (oscillatory, disturbed) flow shear stress activates signaling by PIEZO1 and integrins in endothelial cells / positive regulation of insulin secretion involved in cellular response to glucose stimulus / adenylate cyclase activator activity / trans-Golgi network membrane / GTPase activator activity / TCF dependent signaling in response to WNT / PDZ domain binding / Asymmetric localization of PCP proteins / neuropeptide signaling pathway / negative regulation of inflammatory response to antigenic stimulus / response to prostaglandin E / negative regulation of canonical Wnt signaling pathway / positive regulation of neuron projection development / bone development / G protein-coupled receptor binding / platelet aggregation / G protein-coupled receptor activity / cognition / G-protein beta/gamma-subunit complex binding / neuron differentiation / Olfactory Signaling Pathway / Activation of the phototransduction cascade / blood coagulation / positive regulation of insulin secretion / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / sensory perception of smell / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G beta:gamma signalling through BTK / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
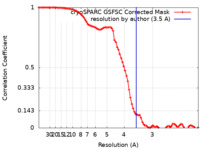
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Lin X / Xu F | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2024 Journal: Cell Discov / Year: 2024Title: A framework for Frizzled-G protein coupling and implications to the PCP signaling pathways. Authors: Zhibin Zhang / Xi Lin / Ling Wei / Yiran Wu / Lu Xu / Lijie Wu / Xiaohu Wei / Suwen Zhao / Xiangjia Zhu / Fei Xu /  Abstract: The ten Frizzled receptors (FZDs) are essential in Wnt signaling and play important roles in embryonic development and tumorigenesis. Among these, FZD6 is closely associated with lens development. ...The ten Frizzled receptors (FZDs) are essential in Wnt signaling and play important roles in embryonic development and tumorigenesis. Among these, FZD6 is closely associated with lens development. Understanding FZD activation mechanism is key to unlock these emerging targets. Here we present the cryo-EM structures of FZD6 and FZD3 which are known to relay non-canonical planar cell polarity (PCP) signaling pathways as well as FZD1 in their G protein-coupled states and in the apo inactive states, respectively. Comparison of the three inactive/active pairs unveiled a shared activation framework among all ten FZDs. Mutagenesis along with imaging and functional analysis on the human lens epithelial tissues suggested potential crosstalk between the G-protein coupling of FZD6 and the PCP signaling pathways. Together, this study provides an integrated understanding of FZD structure and function, and lays the foundation for developing therapeutic modulators to activate or inhibit FZD signaling for a range of disorders including cancers and cataracts. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36111.map.gz emd_36111.map.gz | 49.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36111-v30.xml emd-36111-v30.xml emd-36111.xml emd-36111.xml | 23.4 KB 23.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36111_fsc.xml emd_36111_fsc.xml | 8.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_36111.png emd_36111.png | 33.6 KB | ||
| Filedesc metadata |  emd-36111.cif.gz emd-36111.cif.gz | 7 KB | ||
| Others |  emd_36111_additional_1.map.gz emd_36111_additional_1.map.gz emd_36111_half_map_1.map.gz emd_36111_half_map_1.map.gz emd_36111_half_map_2.map.gz emd_36111_half_map_2.map.gz | 52.3 MB 51.5 MB 51.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36111 http://ftp.pdbj.org/pub/emdb/structures/EMD-36111 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36111 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36111 | HTTPS FTP |
-Related structure data
| Related structure data |  8j9nMC  8j9oC  8jh7C  8jhbC  8jhcC  8jhiC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36111.map.gz / Format: CCP4 / Size: 55.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36111.map.gz / Format: CCP4 / Size: 55.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_36111_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36111_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36111_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FZD1-Gq complex in the ligand-free state
| Entire | Name: FZD1-Gq complex in the ligand-free state |
|---|---|
| Components |
|
-Supramolecule #1: FZD1-Gq complex in the ligand-free state
| Supramolecule | Name: FZD1-Gq complex in the ligand-free state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Frizzled-1
| Macromolecule | Name: Frizzled-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 64.015668 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GVRAQAAGQG PGQGPGPGQQ PPPPPQQQQS GQQYNGERGI SVPDHGYCQP ISIPLCTDIA YNQTIMPNLL GHTNQEDAGL EVHQFYPLV KVQCSAELKF FLCSMYAPVC TVLEQALPPC RSLCERARQG CEALMNKFGF QWPDTLKCEK FPVHGAGELC V GQNTSDKG ...String: GVRAQAAGQG PGQGPGPGQQ PPPPPQQQQS GQQYNGERGI SVPDHGYCQP ISIPLCTDIA YNQTIMPNLL GHTNQEDAGL EVHQFYPLV KVQCSAELKF FLCSMYAPVC TVLEQALPPC RSLCERARQG CEALMNKFGF QWPDTLKCEK FPVHGAGELC V GQNTSDKG TPTPSLLPEF WTSNPQHGGG GHRGGFPGGA GASERGKFSC PRALKVPSYL NYHFLGEKDC GAPCEPTKVY GL MYFGPEE LRFSRTWIGI WSVLCCASTL FTVLTYLVDM RRFSYPERPI IFLSGCYTAV AVAYIAGFLL EDRVVCNDKF AED GARTVA QGTKKEGCTI LFMMLYFFSM ASSIWWVILS LTWFLAAGMK WGHEAIEANS QYFHLAAWAV PAIKTITILA LGQV DGDVL SGVCFVGLNN VDALRGFVLA PLFVYLFIGT SFLLAGFVSL FRIRTIMKHD GTKTEKLEKL MVRIGVFSVL YTVPA TIVI ACYFYEQAFR DQWERSWVAQ SCKSYAIPCP HLQAGGGAPP HPPMSPDFTV FMIKYLMTLI VGITSGFWIW SGKTLN SWR KFYTRLTNSK QGETTV UniProtKB: Frizzled-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(s) subunit alpha isoforms sh...
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short,Guanine nucleotide-binding protein G(q) subunit alpha type: protein_or_peptide / ID: 2 / Details: Chimeric protein MiniGQ / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.8205 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GNSKTEDQRN EEKAQREANK KIEQLRRDKR DARRATHRLL LLGADNSGKS TIVKQMRILH GGSGGSGGTS GIFETKFQVD KVNFHMFDV GGQRDERRKW IQCFNDVTAI IFVVDSSDYN RLQEALNLFK SIWNNRWLRT ISVILFLNKQ DLLAEKVLAG K SKIEDYFP ...String: GNSKTEDQRN EEKAQREANK KIEQLRRDKR DARRATHRLL LLGADNSGKS TIVKQMRILH GGSGGSGGTS GIFETKFQVD KVNFHMFDV GGQRDERRKW IQCFNDVTAI IFVVDSSDYN RLQEALNLFK SIWNNRWLRT ISVILFLNKQ DLLAEKVLAG K SKIEDYFP EFARYTTPED ATPEPGEDPR VTRAKYFIRD EFLRISTASG DGRHYCYPHF TCAVDTENAR RIFNDCKDTI LQ LNLKEYN LV UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short, Guanine nucleotide-binding protein G(q) subunit alpha, Guanine nucleotide-binding protein G(s) subunit alpha isoforms ...UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short, Guanine nucleotide-binding protein G(q) subunit alpha, Guanine nucleotide-binding protein G(s) subunit alpha isoforms short, Guanine nucleotide-binding protein G(s) subunit alpha isoforms short, Guanine nucleotide-binding protein G(q) subunit alpha |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.41693 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPNG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.845078 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFSAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: nanobody Nb35
| Macromolecule | Name: nanobody Nb35 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.054232 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYLLPTAAA GLLLLAAQPA MAMQVQLQES GGGLVQPGGS LRLSCAASGF TFSNYKMNWV RQAPGKGLEW VSDISQSGAS ISYTGSVKG RFTISRDNAK NTLYLQMNSL KPEDTAVYYC ARCPAPFTRD CFDVTSTTYA YRGQGTQVTV |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: OTHER |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)