+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dopamine Receptor D1R-Gs-Rotigotine complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Dopamine / Dopamine receptor / GPCR / D1R / Gs / Rotigotine / Parkinson's disease / Restless legs syndrome / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationdopamine neurotransmitter receptor activity, coupled via Gs / dopamine neurotransmitter receptor activity / cerebral cortex GABAergic interneuron migration / Dopamine receptors / regulation of dopamine uptake involved in synaptic transmission / dopamine binding / operant conditioning / phospholipase C-activating dopamine receptor signaling pathway / heterotrimeric G-protein binding / peristalsis ...dopamine neurotransmitter receptor activity, coupled via Gs / dopamine neurotransmitter receptor activity / cerebral cortex GABAergic interneuron migration / Dopamine receptors / regulation of dopamine uptake involved in synaptic transmission / dopamine binding / operant conditioning / phospholipase C-activating dopamine receptor signaling pathway / heterotrimeric G-protein binding / peristalsis / G protein-coupled receptor complex / modification of postsynaptic structure / positive regulation of neuron migration / habituation / grooming behavior / regulation of dopamine metabolic process / astrocyte development / striatum development / : / dentate gyrus development / positive regulation of potassium ion transport / sensitization / maternal behavior / conditioned taste aversion / arrestin family protein binding / non-motile cilium / G protein-coupled dopamine receptor signaling pathway / mating behavior / adult walking behavior / long-term synaptic depression / : / ciliary membrane / temperature homeostasis / dopamine metabolic process / transmission of nerve impulse / G-protein alpha-subunit binding / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / behavioral fear response / neuronal action potential / adenylate cyclase-activating adrenergic receptor signaling pathway / positive regulation of synaptic transmission, glutamatergic / behavioral response to cocaine / synapse assembly / presynaptic modulation of chemical synaptic transmission / positive regulation of release of sequestered calcium ion into cytosol / response to amphetamine / synaptic transmission, glutamatergic / visual learning / vasodilation / G protein-coupled receptor activity / GABA-ergic synapse / memory / Olfactory Signaling Pathway / Activation of the phototransduction cascade / protein import into nucleus / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / long-term synaptic potentiation / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / G alpha (12/13) signalling events / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / fibroblast proliferation / presynaptic membrane / Ca2+ pathway / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Xu P / Huang S / Zhuang Y / Mao C / Zhang Y / Wang Y / Li H / Jiang Y / Xu HE | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2023 Journal: Cell Res / Year: 2023Title: Structural genomics of the human dopamine receptor system. Authors: Peiyu Xu / Sijie Huang / Brian E Krumm / Youwen Zhuang / Chunyou Mao / Yumu Zhang / Yue Wang / Xi-Ping Huang / Yong-Feng Liu / Xinheng He / Huadong Li / Wanchao Yin / Yi Jiang / Yan Zhang / ...Authors: Peiyu Xu / Sijie Huang / Brian E Krumm / Youwen Zhuang / Chunyou Mao / Yumu Zhang / Yue Wang / Xi-Ping Huang / Yong-Feng Liu / Xinheng He / Huadong Li / Wanchao Yin / Yi Jiang / Yan Zhang / Bryan L Roth / H Eric Xu /   Abstract: The dopaminergic system, including five dopamine receptors (D1R to D5R), plays essential roles in the central nervous system (CNS); and ligands that activate dopamine receptors have been used to ...The dopaminergic system, including five dopamine receptors (D1R to D5R), plays essential roles in the central nervous system (CNS); and ligands that activate dopamine receptors have been used to treat many neuropsychiatric disorders, including Parkinson's Disease (PD) and schizophrenia. Here, we report cryo-EM structures of all five subtypes of human dopamine receptors in complex with G protein and bound to the pan-agonist, rotigotine, which is used to treat PD and restless legs syndrome. The structures reveal the basis of rotigotine recognition in different dopamine receptors. Structural analysis together with functional assays illuminate determinants of ligand polypharmacology and selectivity. The structures also uncover the mechanisms of dopamine receptor activation, unique structural features among the five receptor subtypes, and the basis of G protein coupling specificity. Our work provides a comprehensive set of structural templates for the rational design of specific ligands to treat CNS diseases targeting the dopaminergic system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35683.map.gz emd_35683.map.gz | 55.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35683-v30.xml emd-35683-v30.xml emd-35683.xml emd-35683.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35683.png emd_35683.png | 85.2 KB | ||
| Filedesc metadata |  emd-35683.cif.gz emd-35683.cif.gz | 6.6 KB | ||
| Others |  emd_35683_half_map_1.map.gz emd_35683_half_map_1.map.gz emd_35683_half_map_2.map.gz emd_35683_half_map_2.map.gz | 24.3 MB 24.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35683 http://ftp.pdbj.org/pub/emdb/structures/EMD-35683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35683 | HTTPS FTP |
-Related structure data
| Related structure data |  8irrMC  8irsC  8irtC  8iruC  8irvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35683.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35683.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_35683_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_35683_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Dopamine Receptor D1R-Gs-Rotigotine complex
| Entire | Name: Dopamine Receptor D1R-Gs-Rotigotine complex |
|---|---|
| Components |
|
-Supramolecule #1: Dopamine Receptor D1R-Gs-Rotigotine complex
| Supramolecule | Name: Dopamine Receptor D1R-Gs-Rotigotine complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 29.048932 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCLGNSKTE DQRNEEKAQR EANKMIEKQL QKDKQVYRAT HRLLLLGADN SGKSTIVKQM RIYHVNSGIF ETKFQVDKVN FHMFDVGAQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA ...String: MGCLGNSKTE DQRNEEKAQR EANKMIEKQL QKDKQVYRAT HRLLLLGADN SGKSTIVKQM RIYHVNSGIF ETKFQVDKVN FHMFDVGAQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA RYTTPEDATP EPGEDPRVTR AKYFIRDEFL RISTASGDGR HYCYPHFTCS VDTENARRIF NDCRDIIQRM HL RQYELL |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.020664 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HHHHHHHHMG SLLQSELDQL RQEAEQLKNQ IRDARKACAD ATLSQITNNI DPVGRIQMRT RRTLRGHLAK IYAMHWGTDS RLLVSASQD GKLIIWDSYT TNKVHAIPLR SSWVMTCAYA PSGNYVACGG LDNICSIYNL KTREGNVRVS RELAGHTGYL S CCRFLDDN ...String: HHHHHHHHMG SLLQSELDQL RQEAEQLKNQ IRDARKACAD ATLSQITNNI DPVGRIQMRT RRTLRGHLAK IYAMHWGTDS RLLVSASQD GKLIIWDSYT TNKVHAIPLR SSWVMTCAYA PSGNYVACGG LDNICSIYNL KTREGNVRVS RELAGHTGYL S CCRFLDDN QIVTSSGDTT CALWDIETGQ QTTTFTGHTG DVMSLSLAPD TRLFVSGACD ASAKLWDVRE GMCRQTFTGH ES DINAICF FPNGNAFATG SDDATCRLFD LRADQELMTY SHDNIICGIT SVSFSKSGRL LLAGYDDFNC NVWDALKADR AGV LAGHDN RVSCLGVTDD GMAVATGSWD SFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.432554 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ASNNTASIAQ ARKLVEQLKM EANIDRIKVS KAAADLMAYC EAHAKEDPLL TPVPASENPF REKKFFC UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Nanoboy 35
| Macromolecule | Name: Nanoboy 35 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.845516 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQVQLQESGG GLVQPGGSLR LSCAASGFTF SNYKMNWVRQ APGKGLEWVS DISQSGASIS YTGSVKGRFT ISRDNAKNTL YLQMNSLKP EDTAVYYCAR CPAPFTRDCF DVTSTTYAYR GQGTQVTVSS HHHHHH |
-Macromolecule #5: D(1A) dopamine receptor
| Macromolecule | Name: D(1A) dopamine receptor / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.668707 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DYKDDDDVDM GQPGNGSAFL LAPNGSHAPD HDVTQQRDEE NLYFQGASMR TLNTSAMDGT GLVVERDFSV RILTACFLSL LILSTLLGN TLVCAAVIRF RHLRSKVTNF FVISLAVSDL LVAVLVMPWK AVAEIAGFWP FGSFCNIWVA FDIMCSTASI L NLCVISVD ...String: DYKDDDDVDM GQPGNGSAFL LAPNGSHAPD HDVTQQRDEE NLYFQGASMR TLNTSAMDGT GLVVERDFSV RILTACFLSL LILSTLLGN TLVCAAVIRF RHLRSKVTNF FVISLAVSDL LVAVLVMPWK AVAEIAGFWP FGSFCNIWVA FDIMCSTASI L NLCVISVD RYWAISSPFR YERKMTPKAA FILISVAWTL SVLISFIPVQ LSWHKAKPTS PSDGNATSLA ETIDNCDSSL SR TYAISSS VISFYIPVAI MIVTYTRIYR IAQKQIRRIA ALERAAVHAK NCQTTTGNGK PVECSQPESS FKMSFKRETK VLK TLSVIM GVFVCCWLPF FILNCILPFC GSGETQPFCI DSNTFDVFVW FGWANSSLNP IIYAFNADFR KAFSTLLGCY RLCP ATNNA IETVSINNNG AAMFSSHHEP RGSISKECNL VYLIPHAVGS SEDLKKEEAA GIARPLEKLS PALSVILDYD TDVSL EKIQ PITQNGQHPT HHHHHHHH UniProtKB: D(1A) dopamine receptor |
-Macromolecule #6: Rotigotine
| Macromolecule | Name: Rotigotine / type: ligand / ID: 6 / Number of copies: 1 / Formula: R5F |
|---|---|
| Molecular weight | Theoretical: 315.473 Da |
| Chemical component information | 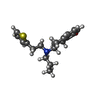 ChemComp-R5F: |
-Macromolecule #7: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 7 / Number of copies: 3 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #8: water
| Macromolecule | Name: water / type: ligand / ID: 8 / Number of copies: 2 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.3 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 6301 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 448516 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)