+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the nucleosome in complex with p53 | |||||||||||||||||||||||||||||||||
 Map data Map data | Cryo-EM structure of the nucleosome in complex with p53 | |||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||
 Keywords Keywords | Transcription factor / Tumor-suppressor / GENE REGULATION-DNA COMPLEX | |||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of helicase activity / Loss of function of TP53 in cancer due to loss of tetramerization ability / Regulation of TP53 Expression / signal transduction by p53 class mediator / negative regulation of G1 to G0 transition / negative regulation of glucose catabolic process to lactate via pyruvate / Transcriptional activation of cell cycle inhibitor p21 / regulation of intrinsic apoptotic signaling pathway by p53 class mediator / negative regulation of pentose-phosphate shunt / ATP-dependent DNA/DNA annealing activity ...negative regulation of helicase activity / Loss of function of TP53 in cancer due to loss of tetramerization ability / Regulation of TP53 Expression / signal transduction by p53 class mediator / negative regulation of G1 to G0 transition / negative regulation of glucose catabolic process to lactate via pyruvate / Transcriptional activation of cell cycle inhibitor p21 / regulation of intrinsic apoptotic signaling pathway by p53 class mediator / negative regulation of pentose-phosphate shunt / ATP-dependent DNA/DNA annealing activity / Activation of NOXA and translocation to mitochondria / regulation of cell cycle G2/M phase transition / oligodendrocyte apoptotic process / negative regulation of miRNA processing / intrinsic apoptotic signaling pathway in response to hypoxia / positive regulation of thymocyte apoptotic process / oxidative stress-induced premature senescence / regulation of tissue remodeling / positive regulation of mitochondrial membrane permeability / regulation of fibroblast apoptotic process / mRNA transcription / bone marrow development / circadian behavior / positive regulation of programmed necrotic cell death / T cell proliferation involved in immune response / regulation of mitochondrial membrane permeability involved in apoptotic process / histone deacetylase regulator activity / RUNX3 regulates CDKN1A transcription / germ cell nucleus / homolactic fermentation / TP53 Regulates Transcription of Death Receptors and Ligands / TP53 regulates transcription of additional cell cycle genes whose exact role in the p53 pathway remain uncertain / Activation of PUMA and translocation to mitochondria / Regulation of TP53 Activity through Association with Co-factors / regulation of DNA damage response, signal transduction by p53 class mediator / negative regulation of glial cell proliferation / negative regulation of neuroblast proliferation / mitochondrial DNA repair / T cell lineage commitment / Formation of Senescence-Associated Heterochromatin Foci (SAHF) / thymocyte apoptotic process / ER overload response / TP53 Regulates Transcription of Caspase Activators and Caspases / cardiac septum morphogenesis / entrainment of circadian clock by photoperiod / B cell lineage commitment / negative regulation of mitophagy / negative regulation of DNA replication / Zygotic genome activation (ZGA) / Association of TriC/CCT with target proteins during biosynthesis / PI5P Regulates TP53 Acetylation / TP53 Regulates Transcription of Genes Involved in Cytochrome C Release / necroptotic process / positive regulation of release of cytochrome c from mitochondria / negative regulation of telomere maintenance via telomerase / SUMOylation of transcription factors / TP53 regulates transcription of several additional cell death genes whose specific roles in p53-dependent apoptosis remain uncertain / TFIID-class transcription factor complex binding / cellular response to actinomycin D / intrinsic apoptotic signaling pathway by p53 class mediator / negative regulation of reactive oxygen species metabolic process / rRNA transcription / Transcriptional Regulation by VENTX / cellular response to UV-C / viral process / general transcription initiation factor binding / replicative senescence / intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress / positive regulation of RNA polymerase II transcription preinitiation complex assembly / neuroblast proliferation / intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / Pyroptosis / positive regulation of execution phase of apoptosis / embryonic organ development / hematopoietic stem cell differentiation / chromosome organization / type II interferon-mediated signaling pathway / response to X-ray / negative regulation of tumor necrosis factor-mediated signaling pathway / TP53 Regulates Transcription of Genes Involved in G1 Cell Cycle Arrest / somitogenesis / hematopoietic progenitor cell differentiation / positive regulation of cardiac muscle cell apoptotic process / core promoter sequence-specific DNA binding / glial cell proliferation / negative regulation of stem cell proliferation / cellular response to glucose starvation / mitophagy / negative regulation of fibroblast proliferation / cis-regulatory region sequence-specific DNA binding / negative regulation of megakaryocyte differentiation / Regulation of TP53 Activity through Acetylation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / positive regulation of intrinsic apoptotic signaling pathway / Replacement of protamines by nucleosomes in the male pronucleus / 14-3-3 protein binding / negative regulation of proteolysis / response to salt stress / CENP-A containing nucleosome Similarity search - Function | |||||||||||||||||||||||||||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||||||||||||||||||||||||||
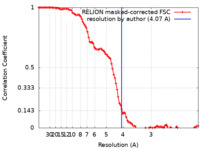
| Method | single particle reconstruction / cryo EM / Resolution: 4.07 Å | |||||||||||||||||||||||||||||||||
 Authors Authors | Nishimura M / Nozawa K / Takizawa Y / Kurumizaka H | |||||||||||||||||||||||||||||||||
| Funding support |  Japan, 10 items Japan, 10 items
| |||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: PNAS Nexus / Year: 2022 Journal: PNAS Nexus / Year: 2022Title: Structural basis for p53 binding to its nucleosomal target DNA sequence. Authors: Masahiro Nishimura / Yoshimasa Takizawa / Kayo Nozawa / Hitoshi Kurumizaka /  Abstract: The tumor suppressor p53 functions as a pioneer transcription factor that binds a nucleosomal target DNA sequence. However, the mechanism by which p53 binds to its target DNA in the nucleosome ...The tumor suppressor p53 functions as a pioneer transcription factor that binds a nucleosomal target DNA sequence. However, the mechanism by which p53 binds to its target DNA in the nucleosome remains elusive. Here we report the cryo-electron microscopy structures of the p53 DNA-binding domain and the full-length p53 protein complexed with a nucleosome containing the 20 base-pair target DNA sequence of p53 (p53BS). In the p53-nucleosome structures, the p53 DNA-binding domain forms a tetramer and specifically binds to the p53BS DNA, located near the entry/exit region of the nucleosome. The nucleosomal position of the p53BS DNA is within the genomic p21 promoter region. The p53 binding peels the DNA from the histone surface, and drastically changes the DNA path around the p53BS on the nucleosome. The C-terminal domain of p53 also binds to the DNA around the center and linker DNA regions of the nucleosome, as revealed by hydroxyl radical footprinting. These results provide important structural information for understanding the mechanism by which p53 binds the nucleosome and changes the chromatin structure for gene activation. | |||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33535.map.gz emd_33535.map.gz | 5.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33535-v30.xml emd-33535-v30.xml emd-33535.xml emd-33535.xml | 26.7 KB 26.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33535_fsc.xml emd_33535_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_33535.png emd_33535.png | 63.6 KB | ||
| Filedesc metadata |  emd-33535.cif.gz emd-33535.cif.gz | 6.9 KB | ||
| Others |  emd_33535_half_map_1.map.gz emd_33535_half_map_1.map.gz emd_33535_half_map_2.map.gz emd_33535_half_map_2.map.gz | 65.3 MB 65.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33535 http://ftp.pdbj.org/pub/emdb/structures/EMD-33535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33535 | HTTPS FTP |
-Related structure data
| Related structure data |  7xzzMC  7xzxC  7xzyC  7y00C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33535.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33535.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the nucleosome in complex with p53 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33535_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33535_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Cryo-EM structure of the nucleosome in complex with p53
+Supramolecule #1: Cryo-EM structure of the nucleosome in complex with p53
+Supramolecule #2: nucleosome, p53
+Supramolecule #3: DNA
+Macromolecule #1: Histone H3.1
+Macromolecule #2: Histone H4
+Macromolecule #3: Histone H2A type 1-B/E
+Macromolecule #4: Histone H2B type 1-J
+Macromolecule #7: Cellular tumor antigen p53
+Macromolecule #5: DNA (169-MER)
+Macromolecule #6: DNA (169-MER)
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)