[English] 日本語
 Yorodumi
Yorodumi- EMDB-31882: The mouse nucleosome structure containing H3mm18 aided by PL2-6 scFv -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31882 | |||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The mouse nucleosome structure containing H3mm18 aided by PL2-6 scFv | |||||||||||||||||||||||||||||||||||||||
 Map data Map data | Nucleosome containing H3mm18 aided by PL2-6 scFv | |||||||||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||||||||
 Keywords Keywords | complex / chromatin / nucleosome / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA complex | |||||||||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationDeposition of new CENPA-containing nucleosomes at the centromere / Inhibition of DNA recombination at telomere / SUMOylation of chromatin organization proteins / DNA Damage/Telomere Stress Induced Senescence / G2/M DNA damage checkpoint / Regulation of endogenous retroelements by KRAB-ZFP proteins / Condensation of Prophase Chromosomes / HDMs demethylate histones / Nonhomologous End-Joining (NHEJ) / Negative Regulation of CDH1 Gene Transcription ...Deposition of new CENPA-containing nucleosomes at the centromere / Inhibition of DNA recombination at telomere / SUMOylation of chromatin organization proteins / DNA Damage/Telomere Stress Induced Senescence / G2/M DNA damage checkpoint / Regulation of endogenous retroelements by KRAB-ZFP proteins / Condensation of Prophase Chromosomes / HDMs demethylate histones / Nonhomologous End-Joining (NHEJ) / Negative Regulation of CDH1 Gene Transcription / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / HDACs deacetylate histones / PRC2 methylates histones and DNA / HATs acetylate histones / Metalloprotease DUBs / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / PKMTs methylate histone lysines / UCH proteinases / Processing of DNA double-strand break ends / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / RMTs methylate histone arginines / Estrogen-dependent gene expression / Ub-specific processing proteases / protein localization to CENP-A containing chromatin / CENP-A containing nucleosome / structural constituent of chromatin / nucleosome / heterochromatin formation / nucleosome assembly / chromatin organization / protein heterodimerization activity / protein-containing complex / DNA binding / nucleoplasm / nucleus Similarity search - Function | |||||||||||||||||||||||||||||||||||||||
| Biological species |  | |||||||||||||||||||||||||||||||||||||||
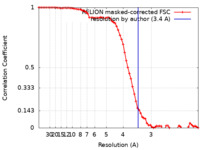
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||||||||||||||||||||||||||||||||
 Authors Authors | Hirai S / Takizawa Y | |||||||||||||||||||||||||||||||||||||||
| Funding support |  Japan, 12 items Japan, 12 items
| |||||||||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2022 Journal: Nucleic Acids Res / Year: 2022Title: Unusual nucleosome formation and transcriptome influence by the histone H3mm18 variant. Authors: Seiya Hirai / Kosuke Tomimatsu / Atsuko Miyawaki-Kuwakado / Yoshimasa Takizawa / Tetsuro Komatsu / Taro Tachibana / Yutaro Fukushima / Yasuko Takeda / Lumi Negishi / Tomoya Kujirai / Masako ...Authors: Seiya Hirai / Kosuke Tomimatsu / Atsuko Miyawaki-Kuwakado / Yoshimasa Takizawa / Tetsuro Komatsu / Taro Tachibana / Yutaro Fukushima / Yasuko Takeda / Lumi Negishi / Tomoya Kujirai / Masako Koyama / Yasuyuki Ohkawa / Hitoshi Kurumizaka /  Abstract: Histone H3mm18 is a non-allelic H3 variant expressed in skeletal muscle and brain in mice. However, its function has remained enigmatic. We found that H3mm18 is incorporated into chromatin in cells ...Histone H3mm18 is a non-allelic H3 variant expressed in skeletal muscle and brain in mice. However, its function has remained enigmatic. We found that H3mm18 is incorporated into chromatin in cells with low efficiency, as compared to H3.3. We determined the structures of the nucleosome core particle (NCP) containing H3mm18 by cryo-electron microscopy, which revealed that the entry/exit DNA regions are drastically disordered in the H3mm18 NCP. Consistently, the H3mm18 NCP is substantially unstable in vitro. The forced expression of H3mm18 in mouse myoblast C2C12 cells markedly suppressed muscle differentiation. A transcriptome analysis revealed that the forced expression of H3mm18 affected the expression of multiple genes, and suppressed a group of genes involved in muscle development. These results suggest a novel gene expression regulation system in which the chromatin landscape is altered by the formation of unusual nucleosomes with a histone variant, H3mm18, and provide important insight into understanding transcription regulation by chromatin. | |||||||||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31882.map.gz emd_31882.map.gz | 2.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31882-v30.xml emd-31882-v30.xml emd-31882.xml emd-31882.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_31882_fsc.xml emd_31882_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_31882.png emd_31882.png | 187.1 KB | ||
| Filedesc metadata |  emd-31882.cif.gz emd-31882.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31882 http://ftp.pdbj.org/pub/emdb/structures/EMD-31882 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31882 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31882 | HTTPS FTP |
-Related structure data
| Related structure data |  7vbmMC  7dbhC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31882.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31882.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Nucleosome containing H3mm18 aided by PL2-6 scFv | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
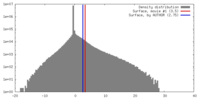
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : The mouse nucleosome containing H3mm18 aided by PL2-6 scFv
| Entire | Name: The mouse nucleosome containing H3mm18 aided by PL2-6 scFv |
|---|---|
| Components |
|
-Supramolecule #1: The mouse nucleosome containing H3mm18 aided by PL2-6 scFv
| Supramolecule | Name: The mouse nucleosome containing H3mm18 aided by PL2-6 scFv type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Histone H3mm18
| Macromolecule | Name: Histone H3mm18 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.350842 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMARTKQT ARKSTGDKAP RKQLATKAAR KSAPSTGGVK KPHCYRPGTV ALREIRSYQK SSELLIRKLP FQRLVLEIAQ DFKTDLCFQ SAAIGALQEA SEAYLVGLFE DTNLCAIHAK RVTIMPKDTQ LAGYICRECA |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.676703 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMSGRGKG GKGLGKGGAK RHRKVLRDNI QGITKPAIRR LARRGGVKRI SGLIYEETRG VLKVFLENVI RDAVTYTEHA KRKTVTAMD VVYALKRQGR TLYGFGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A type 1-B
| Macromolecule | Name: Histone H2A type 1-B / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.447825 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMSGRGKQ GGKARAKAKT RSSRAGLQFP VGRVHRLLRK GNYSERVGAG APVYLAAVLE YLTAEILELA GNAARDNKKT RIIPRHLQL AIRNDEELNK LLGRVTIAQG GVLPNIQAVL LPKKTESHHK AKGK UniProtKB: Histone H2A type 1-B |
-Macromolecule #4: Histone H2B type 3-A
| Macromolecule | Name: Histone H2B type 3-A / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.307559 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMPEPSRS TPAPKKGSKK AITKAQKKDG KKRKRGRKES YSIYVYKVLK QVHPDTGISS KAMGIMNSFV NDIFERIASE ASRLAHYNK RSTITSREVQ TAVRLLLPGE LAKHAVSEGT KAVTKYTSSK UniProtKB: H2B.U histone 2 |
-Macromolecule #5: DNA (126-MER)
| Macromolecule | Name: DNA (126-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 44.520383 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC) ...String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC)(DT)(DA) (DG)(DC)(DA)(DC)(DC)(DG)(DC)(DT)(DT)(DA) (DA)(DA) (DC)(DG)(DC)(DA)(DC)(DG)(DT) (DA)(DC)(DG)(DC)(DG)(DC)(DT)(DG)(DT)(DC) (DC)(DC)(DC) (DC)(DG)(DC)(DG)(DT)(DT) (DT)(DT)(DA)(DA)(DC)(DC)(DG)(DC)(DC)(DA) (DA)(DG)(DG)(DG) (DG)(DA)(DT)(DT)(DA) (DC)(DT)(DC)(DC)(DC)(DT)(DA)(DG)(DT)(DC) (DT)(DC)(DC)(DA)(DG) (DG)(DC)(DA)(DC) (DG)(DT)(DG)(DT)(DC)(DA)(DG)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DC)(DA) (DT)(DC)(DG) (DA)(DT) |
-Macromolecule #6: DNA (126-MER)
| Macromolecule | Name: DNA (126-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 44.99166 KDa |
| Sequence | String: (DA)(DT)(DC)(DG)(DA)(DT)(DG)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA)(DC)(DA) (DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG)(DG)(DA) (DG) (DT)(DA)(DA)(DT)(DC)(DC) ...String: (DA)(DT)(DC)(DG)(DA)(DT)(DG)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA)(DC)(DA) (DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG)(DG)(DA) (DG) (DT)(DA)(DA)(DT)(DC)(DC)(DC)(DC) (DT)(DT)(DG)(DG)(DC)(DG)(DG)(DT)(DT)(DA) (DA)(DA) (DA)(DC)(DG)(DC)(DG)(DG)(DG) (DG)(DG)(DA)(DC)(DA)(DG)(DC)(DG)(DC)(DG) (DT)(DA)(DC) (DG)(DT)(DG)(DC)(DG)(DT) (DT)(DT)(DA)(DA)(DG)(DC)(DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG) (DT)(DC)(DT)(DA)(DC)(DG)(DA)(DC)(DC)(DA) (DA)(DT)(DT)(DG)(DA) (DG)(DC)(DG)(DG) (DC)(DC)(DT)(DC)(DG)(DG)(DC)(DA)(DC)(DC) (DG)(DG)(DG)(DA)(DT)(DT) (DC)(DT)(DG) (DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 58.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)