[English] 日本語
 Yorodumi
Yorodumi- EMDB-26926: Staphylococcus epidermidis RP62A CRISPR effector complex with non... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Staphylococcus epidermidis RP62A CRISPR effector complex with non-self target RNA 1 | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |   Staphylococcus epidermidis RP62A (bacteria) Staphylococcus epidermidis RP62A (bacteria) | |||||||||||||||
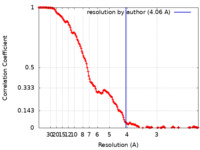
| Method | single particle reconstruction / cryo EM / Resolution: 4.06 Å | |||||||||||||||
 Authors Authors | Smith EM / Ferrell SH / Tokars VL / Mondragon A | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Structures of an active type III-A CRISPR effector complex. Authors: Eric M Smith / Sé Ferrell / Valerie L Tokars / Alfonso Mondragón /  Abstract: Clustered regularly interspaced short palindromic repeats (CRISPR) and their CRISPR-associated proteins (Cas) provide many prokaryotes with an adaptive immune system against invading genetic material. ...Clustered regularly interspaced short palindromic repeats (CRISPR) and their CRISPR-associated proteins (Cas) provide many prokaryotes with an adaptive immune system against invading genetic material. Type III CRISPR systems are unique in that they can degrade both RNA and DNA. In response to invading nucleic acids, they produce cyclic oligoadenylates that act as secondary messengers, activating cellular nucleases that aid in the immune response. Here, we present seven single-particle cryo-EM structures of the type III-A Staphylococcus epidermidis CRISPR effector complex. The structures reveal the intact S. epidermidis effector complex in an apo, ATP-bound, cognate target RNA-bound, and non-cognate target RNA-bound states and illustrate how the effector complex binds and presents crRNA. The complexes bound to target RNA capture the type III-A effector complex in a post-RNA cleavage state. The ATP-bound structures give details about how ATP binds to Cas10 to facilitate cyclic oligoadenylate production. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26926.map.gz emd_26926.map.gz | 141.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26926-v30.xml emd-26926-v30.xml emd-26926.xml emd-26926.xml | 21.4 KB 21.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26926_fsc.xml emd_26926_fsc.xml | 14.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_26926.png emd_26926.png | 77.7 KB | ||
| Masks |  emd_26926_msk_1.map emd_26926_msk_1.map | 282.6 MB |  Mask map Mask map | |
| Others |  emd_26926_half_map_1.map.gz emd_26926_half_map_1.map.gz emd_26926_half_map_2.map.gz emd_26926_half_map_2.map.gz | 261.9 MB 261.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26926 http://ftp.pdbj.org/pub/emdb/structures/EMD-26926 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26926 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26926 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26926.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26926.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_26926_msk_1.map emd_26926_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_26926_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_26926_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Staphylococcus epidermidis RP62a CRISPR effector complex with non...
| Entire | Name: Staphylococcus epidermidis RP62a CRISPR effector complex with non-self target RNA |
|---|---|
| Components |
|
-Supramolecule #1: Staphylococcus epidermidis RP62a CRISPR effector complex with non...
| Supramolecule | Name: Staphylococcus epidermidis RP62a CRISPR effector complex with non-self target RNA type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: CRISPR system single-strand-specific deoxyribonuclease Cas10/Csm1...
| Macromolecule | Name: CRISPR system single-strand-specific deoxyribonuclease Cas10/Csm1 (subtype III-A) type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus epidermidis RP62A (bacteria) Staphylococcus epidermidis RP62A (bacteria) |
| Sequence | String: MNKKNILMYG SLLHDIGKII YRSGDHTFSR GTHSKLGHQF LSQFSEFKDN EVLDNVAYHH YKELAKANL DNDNTAYITY IADNIASGID RRDIIEEGDE EYEKQLFNFD KYTPLYSVFN I VNSEKLKQ TNGKFKFSNE SNIEYPKTEN IQYSSGNYTT LMKDMSHDLE ...String: MNKKNILMYG SLLHDIGKII YRSGDHTFSR GTHSKLGHQF LSQFSEFKDN EVLDNVAYHH YKELAKANL DNDNTAYITY IADNIASGID RRDIIEEGDE EYEKQLFNFD KYTPLYSVFN I VNSEKLKQ TNGKFKFSNE SNIEYPKTEN IQYSSGNYTT LMKDMSHDLE HKLSIKEGTF PS LLQWTES LWQYVPSSTN KNQLIDISLY DHSRITCAIA SCIFDYLNEN NIHNYKDELF SKY ENTKSF YQKEAFLLLS MDMSGIQDFI YNISGSKALK SLRSRSFYLE LMLEVIVDQL LERL ELARA NLLYTGGGHA YLLVSNTDKV KKKITQFNNE LKKWFMSEFT TDLSLSMAFE KCSGD DLMN TSGNYRTIWR NVSSKLSDIK AHKYSAEDIL KLNHFHSYGD RECKECLRSD IDINDD GLC SICEGIINIS NDLRDKSFFV LSETGKLKMP FNKFISVIDY EEAEMLVQNN NQVRIYS KN KPYIGIGIST NLWMCDYDYA SQNQDMREKG IGSYVDREEG VKRLGVVRAD IDNLGATF I SGIPEKYNSI SRTATLSRQL SLFFKYELNH LLENYQITAI YSGGDDLFLI GAWDDIIEA SIYINDKFKE FTLDKLTLSA GVGMFSGKYP VSKMAFETGR LEEAAKTGEK NQISLWLQEK VYNWDEFKK NILEEKLLVL QQGFSQTDEH GKAFIYKMLA LLRNNEAINI ARLAYLLARS K MNEDFTSK IFNWAQNDKD KNQLITALEY YIYQIREAD |
-Macromolecule #2: CRISPR system Cms protein Csm2
| Macromolecule | Name: CRISPR system Cms protein Csm2 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus epidermidis RP62A (bacteria) Staphylococcus epidermidis RP62A (bacteria) |
| Sequence | String: MTFAHEVVKS NVKNVKDRKG KEKQVLFNGL TTSKLRNLME QVNRLYTIAF NSNEDQLNEE FIDELEYLK IKFYYEAGRE KSVDEFLKKT LMFPIIDRVI KKESKKFFLD YCKYFEALVA Y AKYYQKED |
-Macromolecule #3: CRISPR system Cms endoribonuclease Csm3
| Macromolecule | Name: CRISPR system Cms endoribonuclease Csm3 / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus epidermidis RP62A (bacteria) Staphylococcus epidermidis RP62A (bacteria) |
| Sequence | String: MYSKIKISGT IEVVTGLHIG GGGESSMIGA IDSPVVRDLQ TKLPIIPGSS IKGKMRNLLA KHFGLKMKQ ESHNQDDERV LRLFGSSEKG NIQRARLQIS DAFFSEKTKE HFAQNDIAYT E TKFENTIN RLTAVANPRQ IERVTRGSEF DFVFIYNVDE ESQVEDDFEN ...String: MYSKIKISGT IEVVTGLHIG GGGESSMIGA IDSPVVRDLQ TKLPIIPGSS IKGKMRNLLA KHFGLKMKQ ESHNQDDERV LRLFGSSEKG NIQRARLQIS DAFFSEKTKE HFAQNDIAYT E TKFENTIN RLTAVANPRQ IERVTRGSEF DFVFIYNVDE ESQVEDDFEN IEKAIHLLEN DY LGGGGTR GNGRIQFKDT NIETVVGEYD STNLKIK |
-Macromolecule #4: CRISPR system Cms protein Csm4
| Macromolecule | Name: CRISPR system Cms protein Csm4 / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus epidermidis RP62A (bacteria) Staphylococcus epidermidis RP62A (bacteria) |
| Sequence | String: MTLATKVFKL SFKTPVHFGK KRLSDGEMTI TADTLFSALF IETLQLGKDT DWLLNDLIIS DTFPYENEL YYLPKPLIKI DSKEEDNHKA FKKLKYVPVH HYNQYLNGEL SAEDATDLND I FNIGYFSL QTKVSLIAQE TDSSADSEPY SVGTFTFEPE AGLYFIAKGS ...String: MTLATKVFKL SFKTPVHFGK KRLSDGEMTI TADTLFSALF IETLQLGKDT DWLLNDLIIS DTFPYENEL YYLPKPLIKI DSKEEDNHKA FKKLKYVPVH HYNQYLNGEL SAEDATDLND I FNIGYFSL QTKVSLIAQE TDSSADSEPY SVGTFTFEPE AGLYFIAKGS EETLDHLNNI MT ALQYSGL GGKRNAGYGQ FEYEIINNQQ LSKLLNQNGK HSILLSTAMA KKEEIESALK EAR YILTKR SGFVQSTNYS EMLVKKSDFY SFSSGSVFKN IFNGDIFNVG HNGKHPVYRY AKPL WLEV |
-Macromolecule #5: RNA (37-MER)
| Macromolecule | Name: RNA (37-MER) / type: rna / ID: 5 |
|---|---|
| Source (natural) | Organism:  Staphylococcus epidermidis RP62A (bacteria) Staphylococcus epidermidis RP62A (bacteria) |
| Sequence | String: ACGAGAACUA GUAAUAAUUG UCAUUUGCAU ACGUUAC |
-Macromolecule #6: RNA (40-MER)
| Macromolecule | Name: RNA (40-MER) / type: rna / ID: 6 |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: AGCCUGGUAA CGUAUGCAAA UGACAAUUAU UACUAUCCAG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.2 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE Details: cryo-EM grids were prepared by glow discharging for 10 seconds at 15 mA in a Pelco easiGlow glow discharger. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: 3 ul of the protein-RNA complex were added to the grid and allowed to incubate for 10 seconds. After incubation the grid was blotted in a Vitrobot Mark IV (FEI Thermo Fischer) (humidity 95% ...Details: 3 ul of the protein-RNA complex were added to the grid and allowed to incubate for 10 seconds. After incubation the grid was blotted in a Vitrobot Mark IV (FEI Thermo Fischer) (humidity 95% and temperature 4 C) for 3 seconds with a force of 0 before plunge freezing in liquid ethane.. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 2455 / Average electron dose: 49.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 44454 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)