[English] 日本語
 Yorodumi
Yorodumi- EMDB-26331: CryoEM structure of CENP-N promoted nucleosome stacks with CENP-A... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-26331 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of CENP-N promoted nucleosome stacks with CENP-A and palindromic alpha satellite DNA sequence | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | centromere / kinetochore / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationCENP-A containing chromatin assembly / protein localization to chromosome, centromeric region / kinetochore assembly / condensed chromosome, centromeric region / inner kinetochore / establishment of mitotic spindle orientation / mitotic cytokinesis / chromosome, centromeric region / pericentric heterochromatin / negative regulation of megakaryocyte differentiation ...CENP-A containing chromatin assembly / protein localization to chromosome, centromeric region / kinetochore assembly / condensed chromosome, centromeric region / inner kinetochore / establishment of mitotic spindle orientation / mitotic cytokinesis / chromosome, centromeric region / pericentric heterochromatin / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / telomere organization / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / RNA Polymerase I Promoter Opening / Inhibition of DNA recombination at telomere / Assembly of the ORC complex at the origin of replication / Meiotic synapsis / SUMOylation of chromatin organization proteins / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / Resolution of Sister Chromatid Cohesion / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / innate immune response in mucosa / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / chromosome segregation / Transcriptional regulation by small RNAs / RHO GTPases Activate Formins / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / HDMs demethylate histones / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / Negative Regulation of CDH1 Gene Transcription / NoRC negatively regulates rRNA expression / PKMTs methylate histone lysines / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / Pre-NOTCH Transcription and Translation / Meiotic recombination / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Transcriptional regulation of granulopoiesis / Metalloprotease DUBs / RMTs methylate histone arginines / HCMV Early Events / structural constituent of chromatin / Separation of Sister Chromatids / UCH proteinases / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / heterochromatin formation / nucleosome assembly / antibacterial humoral response / E3 ubiquitin ligases ubiquitinate target proteins / HATs acetylate histones / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / chromatin organization / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Processing of DNA double-strand break ends / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / chromatin binding / protein-containing complex / : / DNA binding / RNA binding / extracellular exosome / extracellular region / nucleoplasm / membrane / identical protein binding / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
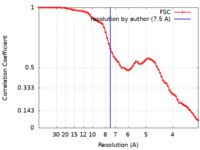
| Method | single particle reconstruction / cryo EM / Resolution: 7.5 Å | |||||||||
 Authors Authors | Zhou K / Luger K | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: CENP-N promotes the compaction of centromeric chromatin. Authors: Keda Zhou / Magdalena Gebala / Dustin Woods / Kousik Sundararajan / Garrett Edwards / Dan Krzizike / Jeff Wereszczynski / Aaron F Straight / Karolin Luger /  Abstract: The histone variant CENP-A is the epigenetic determinant for the centromere, where it is interspersed with canonical H3 to form a specialized chromatin structure that nucleates the kinetochore. How ...The histone variant CENP-A is the epigenetic determinant for the centromere, where it is interspersed with canonical H3 to form a specialized chromatin structure that nucleates the kinetochore. How nucleosomes at the centromere arrange into higher order structures is unknown. Here we demonstrate that the human CENP-A-interacting protein CENP-N promotes the stacking of CENP-A-containing mononucleosomes and nucleosomal arrays through a previously undefined interaction between the α6 helix of CENP-N with the DNA of a neighboring nucleosome. We describe the cryo-EM structures and biophysical characterization of such CENP-N-mediated nucleosome stacks and nucleosomal arrays and demonstrate that this interaction is responsible for the formation of densely packed chromatin at the centromere in the cell. Our results provide first evidence that CENP-A, together with CENP-N, promotes specific chromatin higher order structure at the centromere. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26331.map.gz emd_26331.map.gz | 1.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26331-v30.xml emd-26331-v30.xml emd-26331.xml emd-26331.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26331_fsc.xml emd_26331_fsc.xml | 7.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_26331.png emd_26331.png | 38.3 KB | ||
| Masks |  emd_26331_msk_1.map emd_26331_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-26331.cif.gz emd-26331.cif.gz | 6.3 KB | ||
| Others |  emd_26331_half_map_1.map.gz emd_26331_half_map_1.map.gz emd_26331_half_map_2.map.gz emd_26331_half_map_2.map.gz | 28.4 MB 28.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26331 http://ftp.pdbj.org/pub/emdb/structures/EMD-26331 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26331 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26331 | HTTPS FTP |
-Related structure data
| Related structure data |  7u47MC  7u46C  7u4dC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26331.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26331.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.6422 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_26331_msk_1.map emd_26331_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_26331_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_26331_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : cryoEM structure of centromeric nucleosome in complex with kineto...
| Entire | Name: cryoEM structure of centromeric nucleosome in complex with kinetochore protein CENP-N |
|---|---|
| Components |
|
-Supramolecule #1: cryoEM structure of centromeric nucleosome in complex with kineto...
| Supramolecule | Name: cryoEM structure of centromeric nucleosome in complex with kinetochore protein CENP-N type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Histone H3-like centromeric protein A
| Macromolecule | Name: Histone H3-like centromeric protein A / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 16.02363 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGPRRRSRKP EAPRRRSPSP TPTPGPSRRG PSLGASSHQH SRRRQGWLKE IRKLQKSTHL LIRKLPFSRL AREICVKFTR GVDFNWQAQ ALLALQEAAE AFLVHLFEDA YLLTLHAGRV TLFPKDVQLA RRIRGLEEGL G UniProtKB: Histone H3-like centromeric protein A |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.394426 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A
| Macromolecule | Name: Histone H2A / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.135523 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKQGGK ARAKAKSRSS RAGLQFPVGR VHRLLRKGNY AERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAIR NDEELNKLLG RVTIAQGGVL PNIQAVLLPK KTESHHKAKG K UniProtKB: Histone H2A type 1-C |
-Macromolecule #4: Histone H2B type 1-C/E/F/G/I
| Macromolecule | Name: Histone H2B type 1-C/E/F/G/I / type: protein_or_peptide / ID: 4 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.937213 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSV YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSSK UniProtKB: Histone H2B type 1-C/E/F/G/I |
-Macromolecule #7: Centromere protein N
| Macromolecule | Name: Centromere protein N / type: protein_or_peptide / ID: 7 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 34.870695 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDETVAEFIK RTILKIPMNE LTTILKAWDF LSENQLQTVN FRQRKESVVQ HLIHLCEEKR ASISDAALLD IIYMQFHQHQ KVWDVFQMS KGPGEDVDLF DMKQFKNSFK KILQRALKNV TVSFRETEEN AVWIRIAWGT QYTKPNQYKP TYVVYYSQTP Y AFTSSSML ...String: MDETVAEFIK RTILKIPMNE LTTILKAWDF LSENQLQTVN FRQRKESVVQ HLIHLCEEKR ASISDAALLD IIYMQFHQHQ KVWDVFQMS KGPGEDVDLF DMKQFKNSFK KILQRALKNV TVSFRETEEN AVWIRIAWGT QYTKPNQYKP TYVVYYSQTP Y AFTSSSML RRNTPLLGQA LTIASKHHQI VKMDLRSRYL DSLKAIVFKQ YNQTFETHNS TTPLQERSLG LDINMDSRII HE NIVEKER VQRITQETFG DYPQPQLEFA QYKLETKFKS GLNGSILAER EEHHHHHH UniProtKB: Centromere protein N |
-Macromolecule #5: DNA (147-MER)
| Macromolecule | Name: DNA (147-MER) / type: dna / ID: 5 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 45.368051 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DA)(DT)(DA)(DT)(DC)(DC) (DA)(DC)(DC)(DT)(DG)(DC)(DA)(DG)(DA)(DT) (DA)(DC)(DT)(DA)(DC)(DC)(DA)(DA)(DA) (DA)(DG)(DT)(DG)(DT)(DA)(DT)(DT)(DT)(DG) (DG) (DA)(DA)(DA)(DC)(DT)(DG) ...String: (DA)(DT)(DC)(DA)(DA)(DT)(DA)(DT)(DC)(DC) (DA)(DC)(DC)(DT)(DG)(DC)(DA)(DG)(DA)(DT) (DA)(DC)(DT)(DA)(DC)(DC)(DA)(DA)(DA) (DA)(DG)(DT)(DG)(DT)(DA)(DT)(DT)(DT)(DG) (DG) (DA)(DA)(DA)(DC)(DT)(DG)(DC)(DT) (DC)(DC)(DA)(DT)(DC)(DA)(DA)(DA)(DA)(DG) (DG)(DC) (DA)(DT)(DG)(DT)(DT)(DC)(DA) (DG)(DC)(DT)(DG)(DG)(DA)(DA)(DT)(DC)(DC) (DA)(DG)(DC) (DT)(DG)(DA)(DA)(DC)(DA) (DT)(DG)(DC)(DC)(DT)(DT)(DT)(DT)(DG)(DA) (DT)(DG)(DG)(DA) (DG)(DC)(DA)(DG)(DT) (DT)(DT)(DC)(DC)(DA)(DA)(DA)(DT)(DA)(DC) (DA)(DC)(DT)(DT)(DT) (DT)(DG)(DG)(DT) (DA)(DG)(DT)(DA)(DT)(DC)(DT)(DG)(DC)(DA) (DG)(DG)(DT)(DG)(DG)(DA) (DT)(DA)(DT) (DT)(DG)(DA)(DT) |
-Macromolecule #6: DNA (147-MER)
| Macromolecule | Name: DNA (147-MER) / type: dna / ID: 6 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 45.359035 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DA)(DT)(DA)(DT)(DC)(DC) (DA)(DC)(DC)(DT)(DG)(DC)(DA)(DG)(DA)(DT) (DA)(DC)(DT)(DA)(DC)(DC)(DA)(DA)(DA) (DA)(DG)(DT)(DG)(DT)(DA)(DT)(DT)(DT)(DG) (DG) (DA)(DA)(DA)(DC)(DT)(DG) ...String: (DA)(DT)(DC)(DA)(DA)(DT)(DA)(DT)(DC)(DC) (DA)(DC)(DC)(DT)(DG)(DC)(DA)(DG)(DA)(DT) (DA)(DC)(DT)(DA)(DC)(DC)(DA)(DA)(DA) (DA)(DG)(DT)(DG)(DT)(DA)(DT)(DT)(DT)(DG) (DG) (DA)(DA)(DA)(DC)(DT)(DG)(DC)(DT) (DC)(DC)(DA)(DT)(DC)(DA)(DA)(DA)(DA)(DG) (DG)(DC) (DA)(DT)(DG)(DT)(DT)(DC)(DA) (DG)(DC)(DT)(DG)(DG)(DA)(DT)(DT)(DC)(DC) (DA)(DG)(DC) (DT)(DG)(DA)(DA)(DC)(DA) (DT)(DG)(DC)(DC)(DT)(DT)(DT)(DT)(DG)(DA) (DT)(DG)(DG)(DA) (DG)(DC)(DA)(DG)(DT) (DT)(DT)(DC)(DC)(DA)(DA)(DA)(DT)(DA)(DC) (DA)(DC)(DT)(DT)(DT) (DT)(DG)(DG)(DT) (DA)(DG)(DT)(DA)(DT)(DC)(DT)(DG)(DC)(DA) (DG)(DG)(DT)(DG)(DG)(DA) (DT)(DA)(DT) (DT)(DG)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)