[English] 日本語
 Yorodumi
Yorodumi- EMDB-25559: Cryo-EM structure of the extracellular module of the full-length ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25559 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
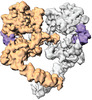
| Title | Cryo-EM structure of the extracellular module of the full-length EGFR L834R bound to EGF. "tips-separated" conformation | |||||||||
 Map data Map data | L834R:EGF_sep | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | receptor tyrosine kinases / epidermal growth factor receptor / SIGNALING PROTEIN / SIGNALING PROTEIN-RECEPTOR complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of hyaluronan biosynthetic process / negative regulation of secretion / negative regulation of cholesterol efflux / positive regulation of cerebellar granule cell precursor proliferation / Developmental Lineage of Multipotent Pancreatic Progenitor Cells / regulation of protein localization to cell surface / positive regulation of protein localization to early endosome / cerebellar granule cell precursor proliferation / regulation of calcium ion import / positive regulation of epithelial tube formation ...positive regulation of hyaluronan biosynthetic process / negative regulation of secretion / negative regulation of cholesterol efflux / positive regulation of cerebellar granule cell precursor proliferation / Developmental Lineage of Multipotent Pancreatic Progenitor Cells / regulation of protein localization to cell surface / positive regulation of protein localization to early endosome / cerebellar granule cell precursor proliferation / regulation of calcium ion import / positive regulation of epithelial tube formation / transmembrane receptor protein tyrosine kinase activator activity / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin / positive regulation of peptidyl-threonine phosphorylation / multivesicular body, internal vesicle lumen / negative regulation of cardiocyte differentiation / Shc-EGFR complex / positive regulation of protein kinase C signaling / regulation of receptor signaling pathway via JAK-STAT / Inhibition of Signaling by Overexpressed EGFR / epidermal growth factor receptor activity / EGFR interacts with phospholipase C-gamma / positive regulation of DNA binding / NFE2L2 regulating tumorigenic genes / positive regulation of ubiquitin-dependent protein catabolic process / epidermal growth factor binding / regulation of peptidyl-tyrosine phosphorylation / epidermal growth factor receptor binding / response to UV-A / ubiquitin-dependent endocytosis / PLCG1 events in ERBB2 signaling / morphogenesis of an epithelial fold / PTK6 promotes HIF1A stabilization / ERBB2 Activates PTK6 Signaling / branching morphogenesis of an epithelial tube / digestive tract morphogenesis / ERBB2-EGFR signaling pathway / Signaling by EGFR / eyelid development in camera-type eye / intracellular vesicle / cerebral cortex cell migration / ERBB2 Regulates Cell Motility / Developmental Lineage of Mammary Gland Myoepithelial Cells / protein insertion into membrane / protein tyrosine kinase activator activity / Signaling by ERBB4 / Respiratory syncytial virus (RSV) attachment and entry / mammary gland alveolus development / negative regulation of epidermal growth factor receptor signaling pathway / positive regulation of receptor internalization / PI3K events in ERBB2 signaling / positive regulation of peptidyl-serine phosphorylation / positive regulation of phosphorylation / hair follicle development / Estrogen-dependent nuclear events downstream of ESR-membrane signaling / MAP kinase kinase kinase activity / embryonic placenta development / GAB1 signalosome / salivary gland morphogenesis / xenobiotic transport / positive regulation of G1/S transition of mitotic cell cycle / positive regulation of epidermal growth factor receptor signaling pathway / ERK1 and ERK2 cascade / Signaling by ERBB2 / TFAP2 (AP-2) family regulates transcription of growth factors and their receptors / positive regulation of endothelial cell proliferation / transmembrane receptor protein tyrosine kinase activity / positive regulation of mitotic nuclear division / GRB2 events in EGFR signaling / SHC1 events in EGFR signaling / EGFR Transactivation by Gastrin / positive regulation of endothelial cell migration / GRB2 events in ERBB2 signaling / epithelial cell proliferation / guanyl-nucleotide exchange factor activity / platelet alpha granule lumen / SHC1 events in ERBB2 signaling / ossification / basal plasma membrane / cellular response to epidermal growth factor stimulus / positive regulation of DNA replication / positive regulation of epithelial cell proliferation / positive regulation of DNA repair / Signal transduction by L1 / positive regulation of protein localization to plasma membrane / growth factor activity / cellular response to estradiol stimulus / NOTCH3 Activation and Transmission of Signal to the Nucleus / cellular response to amino acid stimulus / phosphatidylinositol 3-kinase/protein kinase B signal transduction / sperm end piece / clathrin-coated endocytic vesicle membrane / Signaling by ERBB2 TMD/JMD mutants / EGFR downregulation / receptor protein-tyrosine kinase / Constitutive Signaling by EGFRvIII / cell-cell adhesion / negative regulation of protein catabolic process / Signaling by ERBB2 ECD mutants / Signaling by ERBB2 KD Mutants / positive regulation of protein phosphorylation Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Huang Y / Ognjenovic J | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2021 Journal: Elife / Year: 2021Title: A molecular mechanism for the generation of ligand-dependent differential outputs by the epidermal growth factor receptor. Authors: Yongjian Huang / Jana Ognjenovic / Deepti Karandur / Kate Miller / Alan Merk / Sriram Subramaniam / John Kuriyan /   Abstract: The epidermal growth factor receptor (EGFR) is a receptor tyrosine kinase that couples the binding of extracellular ligands, such as EGF and transforming growth factor-α (TGF-α), to the initiation ...The epidermal growth factor receptor (EGFR) is a receptor tyrosine kinase that couples the binding of extracellular ligands, such as EGF and transforming growth factor-α (TGF-α), to the initiation of intracellular signaling pathways. EGFR binds to EGF and TGF-α with similar affinity, but generates different signals from these ligands. To address the mechanistic basis of this phenomenon, we have carried out cryo-EM analyses of human EGFR bound to EGF and TGF-α. We show that the extracellular module adopts an ensemble of dimeric conformations when bound to either EGF or TGF-α. The two extreme states of this ensemble represent distinct ligand-bound quaternary structures in which the membrane-proximal tips of the extracellular module are either juxtaposed or separated. EGF and TGF-α differ in their ability to maintain the conformation with the membrane-proximal tips of the extracellular module separated, and this conformation is stabilized preferentially by an oncogenic EGFR mutation. Close proximity of the transmembrane helices at the junction with the extracellular module has been associated previously with increased EGFR activity. Our results show how EGFR can couple the binding of different ligands to differential modulation of this proximity, thereby suggesting a molecular mechanism for the generation of ligand-sensitive differential outputs in this receptor family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25559.map.gz emd_25559.map.gz | 62.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25559-v30.xml emd-25559-v30.xml emd-25559.xml emd-25559.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25559.png emd_25559.png | 184.3 KB | ||
| Filedesc metadata |  emd-25559.cif.gz emd-25559.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25559 http://ftp.pdbj.org/pub/emdb/structures/EMD-25559 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25559 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25559 | HTTPS FTP |
-Related structure data
| Related structure data |  7sz1MC  7sydC  7syeC  7sz0C  7sz5C  7sz7C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25559.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25559.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | L834R:EGF_sep | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0794 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : full-length human EGFR(L834R):EGF complex
| Entire | Name: full-length human EGFR(L834R):EGF complex |
|---|---|
| Components |
|
-Supramolecule #1: full-length human EGFR(L834R):EGF complex
| Supramolecule | Name: full-length human EGFR(L834R):EGF complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Macromolecule #1: Epidermal growth factor receptor
| Macromolecule | Name: Epidermal growth factor receptor / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: receptor protein-tyrosine kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 134.477375 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRPSGTAGAA LLALLAALCP ASRALEEKKV CQGTSNKLTQ LGTFEDHFLS LQRMFNNCEV VLGNLEITYV QRNYDLSFLK TIQEVAGYV LIALNTVERI PLENLQIIRG NMYYENSYAL AVLSNYDANK TGLKELPMRN LQEILHGAVR FSNNPALCNV E SIQWRDIV ...String: MRPSGTAGAA LLALLAALCP ASRALEEKKV CQGTSNKLTQ LGTFEDHFLS LQRMFNNCEV VLGNLEITYV QRNYDLSFLK TIQEVAGYV LIALNTVERI PLENLQIIRG NMYYENSYAL AVLSNYDANK TGLKELPMRN LQEILHGAVR FSNNPALCNV E SIQWRDIV SSDFLSNMSM DFQNHLGSCQ KCDPSCPNGS CWGAGEENCQ KLTKIICAQQ CSGRCRGKSP SDCCHNQCAA GC TGPRESD CLVCRKFRNE ATCKDTCPPL MLYNPTTYQM DVNPEGKYSF GATCVKKCPR NYVVTDHGSC VRACGADSYE MEE DGVRKC KKCEGPCRKV CNGIGIGEFK DSLSINATNI KHFKNCTSIS GDLHILPVAF RGDSFTHTPP LDPQELDILK TVKE ITGFL LIQAWPENRT DLHAFENLEI IRGRTKQHGQ FSLAVVSLNI TSLGLRSLKE ISDGDVIISG NKNLCYANTI NWKKL FGTS GQKTKIISNR GENSCKATGQ VCHALCSPEG CWGPEPRDCV SCRNVSRGRE CVDKCNLLEG EPREFVENSE CIQCHP ECL PQAMNITCTG RGPDNCIQCA HYIDGPHCVK TCPAGVMGEN NTLVWKYADA GHVCHLCHPN CTYGCTGPGL EGCPTNG PK IPSIATGMVG ALLLLLVVAL GIGLFMRRRH IVRKRTLRRL LQERELVEPL TPSGEAPNQA LLRILKETEF KKIKVLGS G AFGTVYKGLW IPEGEKVKIP VAIKELREAT SPKANKEILD EAYVMASVDN PHVCRLLGIC LTSTVQLITQ LMPFGCLLD YVREHKDNIG SQYLLNWCVQ IAKGMNYLED RRLVHRDLAA RNVLVKTPQH VKITDFGRAK LLGAEEKEYH AEGGKVPIKW MALESILHR IYTHQSDVWS YGVTVWELMT FGSKPYDGIP ASEISSILEK GERLPQPPIC TIDVYMIMVK CWMIDADSRP K FRELIIEF SKMARDPQRY LVIQGDERMH LPSPTDSNFY RALMDEEDMD DVVDADEYLI PQQGFFSSPS TSRTPLLSSL SA TSNNSTV ACIDRNGLQS CPIKEDSFLQ RYSSDPTGAL TEDSIDDTFL PVPEYINQSV PKRPAGSVQN PVYHNQPLNP APS RDPHYQ DPHSTAVGNP EYLNTVQPTC VNSTFDSPAH WAQKGSHQIS LDNPDYQQDF FPKEAKPNGI FKGSTAENAE YLRV APQSS EFIGA UniProtKB: Epidermal growth factor receptor |
-Macromolecule #2: Epidermal growth factor
| Macromolecule | Name: Epidermal growth factor / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 6.229027 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: NSDSECPLSH DGYCLHDGVC MYIEALDKYA CNCVVGYIGE RCQYRDLKWW ELR UniProtKB: Pro-epidermal growth factor |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















