+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24107 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
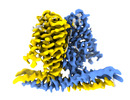
| Title | Cryo-EM structure of TACAN in the apo form (TMEM120A) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | lipid metabolism / coenzyme A / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcoenzyme A binding / protein heterooligomerization / nuclear inner membrane / monoatomic ion channel activity / detection of mechanical stimulus involved in sensory perception of pain / fat cell differentiation / antiviral innate immune response / protein homooligomerization / monoatomic ion transmembrane transport / endoplasmic reticulum ...coenzyme A binding / protein heterooligomerization / nuclear inner membrane / monoatomic ion channel activity / detection of mechanical stimulus involved in sensory perception of pain / fat cell differentiation / antiviral innate immune response / protein homooligomerization / monoatomic ion transmembrane transport / endoplasmic reticulum / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Niu Y / Tao X | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2021 Journal: Elife / Year: 2021Title: Analysis of the mechanosensor channel functionality of TACAN. Authors: Yiming Niu / Xiao Tao / George Vaisey / Paul Dominic B Olinares / Hanan Alwaseem / Brian T Chait / Roderick MacKinnon /  Abstract: Mechanosensitive ion channels mediate transmembrane ion currents activated by mechanical forces. A mechanosensitive ion channel called TACAN was recently reported. We began to study TACAN with the ...Mechanosensitive ion channels mediate transmembrane ion currents activated by mechanical forces. A mechanosensitive ion channel called TACAN was recently reported. We began to study TACAN with the intent to understand how it senses mechanical forces and functions as an ion channel. Using cellular patch-recording methods, we failed to identify mechanosensitive ion channel activity. Using membrane reconstitution methods, we found that TACAN, at high protein concentrations, produces heterogeneous conduction levels that are not mechanosensitive and are most consistent with disruptions of the lipid bilayer. We determined the structure of TACAN using single-particle cryo-electron microscopy and observed that it is a symmetrical dimeric transmembrane protein. Each protomer contains an intracellular-facing cleft with a coenzyme A cofactor, confirmed by mass spectrometry. The TACAN protomer is related in three-dimensional structure to a fatty acid elongase, ELOVL7. Whilst its physiological function remains unclear, we anticipate that TACAN is not a mechanosensitive ion channel. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24107.map.gz emd_24107.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24107-v30.xml emd-24107-v30.xml emd-24107.xml emd-24107.xml | 11.8 KB 11.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24107.png emd_24107.png | 101.5 KB | ||
| Filedesc metadata |  emd-24107.cif.gz emd-24107.cif.gz | 5.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24107 http://ftp.pdbj.org/pub/emdb/structures/EMD-24107 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24107 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24107 | HTTPS FTP |
-Related structure data
| Related structure data |  7n0kMC  7n0lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24107.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24107.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Homo-dimeric assembly of wild type TACAN(TMEM120A)
| Entire | Name: Homo-dimeric assembly of wild type TACAN(TMEM120A) |
|---|---|
| Components |
|
-Supramolecule #1: Homo-dimeric assembly of wild type TACAN(TMEM120A)
| Supramolecule | Name: Homo-dimeric assembly of wild type TACAN(TMEM120A) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ion channel TACAN
| Macromolecule | Name: Ion channel TACAN / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.797297 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MQSPPPDPLG DCLRNWEDLQ QDFQGIQETH RLYRLKLEEL TKLQANCTNS ITRQKKRLQE LALVLKKCRP SLPSESMEAA QELENQMKE RQGLFFDMEA YLPKKNGLYL SLVLGNVNVT LLSKQAKFAY KDEYEKFKLY LTIILIVISF TCRFLLNSRV T DAAFNFLL ...String: MQSPPPDPLG DCLRNWEDLQ QDFQGIQETH RLYRLKLEEL TKLQANCTNS ITRQKKRLQE LALVLKKCRP SLPSESMEAA QELENQMKE RQGLFFDMEA YLPKKNGLYL SLVLGNVNVT LLSKQAKFAY KDEYEKFKLY LTIILIVISF TCRFLLNSRV T DAAFNFLL VWYYCTLTIR ESILINNGSR IKGWWVFHHY VSTFLSGVML TWPDGLMYQK FRNQFLSFSM YQSFVQFLQY YY QSGCLYR LRALGERHTM DLTVEGFQSW MWRGLTFLLP FLFFGHFWQL FNALTLFNLA RDPECKEWQV LMCGFPFLLL FLG NFFTTL RVVHQKFHSQ QHGNKKD UniProtKB: Transmembrane protein 120A |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL |
|---|---|
| Buffer | pH: 7.4 Details: 20 mM HEPES pH 7.4, 250 mM NaCl, and 0.06% Digitonin (w/v) |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number grids imaged: 1 / Average exposure time: 10.0 sec. / Average electron dose: 75.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















