+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23813 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Autoinhibited B-Raf:(14-3-3)2:MEK complex with resolved RBD | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | B-Raf / MEK / 14-3-3 / B-Raf complex / B-Raf monomer / Inactive B-Raf / Serine/threonine-protein kinase B-raf / RBD / signaling protein / SIGNALING PROTEIN-TRANSFERASE complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsynaptic target recognition / negative regulation of homotypic cell-cell adhesion / epithelial cell proliferation involved in lung morphogenesis / Golgi reassembly / positive regulation of endodermal cell differentiation / negative regulation of hypoxia-induced intrinsic apoptotic signaling pathway / regulation of vascular associated smooth muscle contraction / CD4-positive, alpha-beta T cell differentiation / NOTCH4 Activation and Transmission of Signal to the Nucleus / positive regulation of axon regeneration ...synaptic target recognition / negative regulation of homotypic cell-cell adhesion / epithelial cell proliferation involved in lung morphogenesis / Golgi reassembly / positive regulation of endodermal cell differentiation / negative regulation of hypoxia-induced intrinsic apoptotic signaling pathway / regulation of vascular associated smooth muscle contraction / CD4-positive, alpha-beta T cell differentiation / NOTCH4 Activation and Transmission of Signal to the Nucleus / positive regulation of axon regeneration / CD4-positive or CD8-positive, alpha-beta T cell lineage commitment / negative regulation of synaptic vesicle exocytosis / mitogen-activated protein kinase kinase / establishment of Golgi localization / positive regulation of muscle contraction / Golgi inheritance / placenta blood vessel development / MAP kinase scaffold activity / Signalling to p38 via RIT and RIN / regulation of axon regeneration / respiratory system process / cerebellar cortex formation / labyrinthine layer development / head morphogenesis / ARMS-mediated activation / myeloid progenitor cell differentiation / tube formation / melanosome transport / endothelial cell apoptotic process / regulation of synapse maturation / Signaling by MAP2K mutants / SHOC2 M1731 mutant abolishes MRAS complex function / Gain-of-function MRAS complexes activate RAF signaling / negative regulation of fibroblast migration / positive regulation of D-glucose transmembrane transport / Rap1 signalling / establishment of protein localization to membrane / type B pancreatic cell proliferation / central nervous system neuron differentiation / vesicle transport along microtubule / positive regulation of Ras protein signal transduction / regulation of Golgi inheritance / mitogen-activated protein kinase kinase kinase binding / regulation of T cell differentiation / positive regulation of axonogenesis / trachea formation / KSRP (KHSRP) binds and destabilizes mRNA / negative regulation of protein localization to nucleus / triglyceride homeostasis / regulation of early endosome to late endosome transport / Negative feedback regulation of MAPK pathway / regulation of stress-activated MAPK cascade / Frs2-mediated activation / GP1b-IX-V activation signalling / stress fiber assembly / MAPK3 (ERK1) activation / ERBB2-ERBB3 signaling pathway / face development / MAP kinase kinase activity / endodermal cell differentiation / regulation of neurotransmitter receptor localization to postsynaptic specialization membrane / Bergmann glial cell differentiation / positive regulation of protein serine/threonine kinase activity / thyroid gland development / positive regulation of ATP biosynthetic process / Regulation of localization of FOXO transcription factors / Interleukin-3, Interleukin-5 and GM-CSF signaling / Uptake and function of anthrax toxins / somatic stem cell population maintenance / positive regulation of peptidyl-serine phosphorylation / synaptic vesicle exocytosis / Activation of BAD and translocation to mitochondria / phosphoserine residue binding / MAP kinase kinase kinase activity / protein kinase activator activity / negative regulation of endothelial cell apoptotic process / regulation of ERK1 and ERK2 cascade / SARS-CoV-2 targets host intracellular signalling and regulatory pathways / response to axon injury / Schwann cell development / postsynaptic modulation of chemical synaptic transmission / protein targeting / cellular response to glucose starvation / keratinocyte differentiation / SARS-CoV-1 targets host intracellular signalling and regulatory pathways / RHO GTPases activate PKNs / Chk1/Chk2(Cds1) mediated inactivation of Cyclin B:Cdk1 complex / ERK1 and ERK2 cascade / positive regulation of stress fiber assembly / neuron projection morphogenesis / negative regulation of TORC1 signaling / myelination / positive regulation of substrate adhesion-dependent cell spreading / protein serine/threonine/tyrosine kinase activity / Transcriptional and post-translational regulation of MITF-M expression and activity / substrate adhesion-dependent cell spreading / lung development / positive regulation of autophagy / insulin-like growth factor receptor signaling pathway / cellular response to calcium ion Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.66 Å | |||||||||
 Authors Authors | Martinez Fiesco JA / Ping Z | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural insights into the BRAF monomer-to-dimer transition mediated by RAS binding. Authors: Juliana A Martinez Fiesco / David E Durrant / Deborah K Morrison / Ping Zhang /  Abstract: RAF kinases are essential effectors of RAS, but how RAS binding initiates the conformational changes needed for autoinhibited RAF monomers to form active dimers has remained unclear. Here, we present ...RAF kinases are essential effectors of RAS, but how RAS binding initiates the conformational changes needed for autoinhibited RAF monomers to form active dimers has remained unclear. Here, we present cryo-electron microscopy structures of full-length BRAF complexes derived from mammalian cells: autoinhibited, monomeric BRAF:14-3-3:MEK and BRAF:14-3-3 complexes, and an inhibitor-bound, dimeric BRAF:14-3-3 complex, at 3.7, 4.1, and 3.9 Å resolution, respectively. In both autoinhibited, monomeric structures, the RAS binding domain (RBD) of BRAF is resolved, revealing that the RBD forms an extensive contact interface with the 14-3-3 protomer bound to the BRAF C-terminal site and that key basic residues required for RBD-RAS binding are exposed. Moreover, through structure-guided mutational studies, our findings indicate that RAS-RAF binding is a dynamic process and that RBD residues at the center of the RBD:14-3-3 interface have a dual function, first contributing to RAF autoinhibition and then to the full spectrum of RAS-RBD interactions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23813.map.gz emd_23813.map.gz | 2.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23813-v30.xml emd-23813-v30.xml emd-23813.xml emd-23813.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
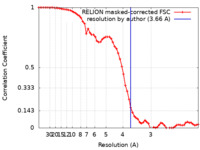
| FSC (resolution estimation) |  emd_23813_fsc.xml emd_23813_fsc.xml | 6.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_23813.png emd_23813.png | 52 KB | ||
| Filedesc metadata |  emd-23813.cif.gz emd-23813.cif.gz | 7.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23813 http://ftp.pdbj.org/pub/emdb/structures/EMD-23813 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23813 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23813 | HTTPS FTP |
-Related structure data
| Related structure data |  7mfdMC  7mfeC  7mffC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23813.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23813.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.058 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Autoinhibited B-Raf:(14-3-3)2:MEK complex with resolved RBD
| Entire | Name: Autoinhibited B-Raf:(14-3-3)2:MEK complex with resolved RBD |
|---|---|
| Components |
|
-Supramolecule #1: Autoinhibited B-Raf:(14-3-3)2:MEK complex with resolved RBD
| Supramolecule | Name: Autoinhibited B-Raf:(14-3-3)2:MEK complex with resolved RBD type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|
-Supramolecule #2: B-raf
| Supramolecule | Name: B-raf / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: MEK
| Supramolecule | Name: MEK / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: 14-3-3 protein zeta/delta
| Supramolecule | Name: 14-3-3 protein zeta/delta / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Serine/threonine-protein kinase B-raf
| Macromolecule | Name: Serine/threonine-protein kinase B-raf / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 84.697695 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAALSGGGGG GAEPGQALFN GDMEPEAGAG AGAAASSAAD PAIPEEVWNI KQMIKLTQEH IEALLDKFGG EHNPPSIYLE AYEEYTSKL DALQQREQQL LESLGNGTDF SVSSSASMDT VTSSSSSSLS VLPSSLSVFQ NPTDVARSNP KSPQKPIVRV F LPNKQRTV ...String: MAALSGGGGG GAEPGQALFN GDMEPEAGAG AGAAASSAAD PAIPEEVWNI KQMIKLTQEH IEALLDKFGG EHNPPSIYLE AYEEYTSKL DALQQREQQL LESLGNGTDF SVSSSASMDT VTSSSSSSLS VLPSSLSVFQ NPTDVARSNP KSPQKPIVRV F LPNKQRTV VPARCGVTVR DSLKKALMMR GLIPECCAVY RIQDGEKKPI GWDTDISWLT GEELHVEVLE NVPLTTHNFV RK TFFTLAF CDFCRKLLFQ GFRCQTCGYK FHQRCSTEVP LMCVNYDQLD LLFVSKFFEH HPIPQEEASL AETALTSGSS PSA PASDSI GPQILTSPSP SKSIPIPQPF RPADEDHRNQ FGQRDRSS(SEP)A PNVHINTIEP VNIDDLIRDQ GFRGDGGSTT GLSATPPAS LPGSLTNVKA LQKSPGPQRE RKSSSSSEDR NRMKTLGRRD SSDDWEIPDG QITVGQRIGS GSFGTVYKGK W HGDVAVKM LNVTAPTPQQ LQAFKNEVGV LRKTRHVNIL LFMGYSTKPQ LAIVTQWCEG SSLYHHLHII ETKFEMIKLI DI ARQTAQG MDYLHAKSII HRDLKSNNIF LHEDLTVKIG DFGLATVKSR WSGSHQFEQL SGSILWMAPE VIRMQDKNPY SFQ SDVYAF GIVLYELMTG QLPYSNINNR DQIIFMVGRG YLSPDLSKVR SNCPKAMKRL MAECLKKKRD ERPLFPQILA SIEL LARSL PKIHRSA(SEP)EP SLNRAGFQTE DFSLYACASP KTPIQAGGYG AFPVH UniProtKB: Serine/threonine-protein kinase B-raf |
-Macromolecule #2: Dual specificity mitogen-activated protein kinase kinase 1
| Macromolecule | Name: Dual specificity mitogen-activated protein kinase kinase 1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: mitogen-activated protein kinase kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.493938 KDa |
| Sequence | String: MPKKKPTPIQ LNPAPDGSAV NGTSSAETNL EALQKKLEEL ELDEQQRKRL EAFLTQKQKV GELKDDDFEK ISELGAGNGG VVFKVSHKP SGLVMARKLI HLEIKPAIRN QIIRELQVLH ECNSPYIVGF YGAFYSDGEI SICMEHMDGG SLDQVLKKAG R IPEQILGK ...String: MPKKKPTPIQ LNPAPDGSAV NGTSSAETNL EALQKKLEEL ELDEQQRKRL EAFLTQKQKV GELKDDDFEK ISELGAGNGG VVFKVSHKP SGLVMARKLI HLEIKPAIRN QIIRELQVLH ECNSPYIVGF YGAFYSDGEI SICMEHMDGG SLDQVLKKAG R IPEQILGK VSIAVIKGLT YLREKHKIMH RDVKPSNILV NSRGEIKLCD FGVSGQLIDS MANSFVGTRS YMSPERLQGT HY SVQSDIW SMGLSLVEMA VGRYPIPPPD AKELELMFGC QVEGDAAETP PRPRTPGRPL SSYGMDSRPP MAIFELLDYI VNE PPPKLP SGVFSLEFQD FVNKCLIKNP AERADLKQLM VHAFIKRSDA EEVDFAGWLC STIGLNQPST PTHAAGV UniProtKB: Dual specificity mitogen-activated protein kinase kinase 1 |
-Macromolecule #3: 14-3-3 protein zeta/delta
| Macromolecule | Name: 14-3-3 protein zeta/delta / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.777092 KDa |
| Sequence | String: MDKNELVQKA KLAEQAERYD DMAACMKSVT EQGAELSNEE RNLLSVAYKN VVGARRSSWR VVSSIEQKTE GAEKKQQMAR EYREKIETE LRDICNDVLS LLEKFLIPNA SQAESKVFYL KMKGDYYRYL AEVAAGDDKK GIVDQSQQAY QEAFEISKKE M QPTHPIRL ...String: MDKNELVQKA KLAEQAERYD DMAACMKSVT EQGAELSNEE RNLLSVAYKN VVGARRSSWR VVSSIEQKTE GAEKKQQMAR EYREKIETE LRDICNDVLS LLEKFLIPNA SQAESKVFYL KMKGDYYRYL AEVAAGDDKK GIVDQSQQAY QEAFEISKKE M QPTHPIRL GLALNFSVFY YEILNSPEKA CSLAKTAFDE AIAELDTLSE ESYKDSTLIM QLLRDNLTLW TSDTQGDEAE AG EGGEN UniProtKB: 14-3-3 protein zeta/delta |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #5: N-(3-fluoro-4-{[4-methyl-2-oxo-7-(pyrimidin-2-yloxy)-2H-chromen-3...
| Macromolecule | Name: N-(3-fluoro-4-{[4-methyl-2-oxo-7-(pyrimidin-2-yloxy)-2H-chromen-3-yl]methyl}pyridin-2-yl)-N'-methylsulfuric diamide type: ligand / ID: 5 / Number of copies: 1 / Formula: CHU |
|---|---|
| Molecular weight | Theoretical: 471.462 Da |
| Chemical component information |  ChemComp-CHU: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Details: 20mAmp | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 57.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)