[English] 日本語
 Yorodumi
Yorodumi- EMDB-23588: Cryo-EM map of the elongation module of the bacillamide NRPS, Bmd... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23588 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC | |||||||||
 Map data Map data | EM Map of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Thermoactinomyces vulgaris (bacteria) Thermoactinomyces vulgaris (bacteria) | |||||||||
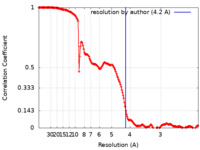
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Sharon I / Strauss M / Schmeing TM | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structures and function of a tailoring oxidase in complex with a nonribosomal peptide synthetase module. Authors: Camille Marie Fortinez / Kristjan Bloudoff / Connor Harrigan / Itai Sharon / Mike Strauss / T Martin Schmeing /  Abstract: Nonribosomal peptide synthetases (NRPSs) are large modular enzymes that synthesize secondary metabolites and natural product therapeutics. Most NRPS biosynthetic pathways include an NRPS and ...Nonribosomal peptide synthetases (NRPSs) are large modular enzymes that synthesize secondary metabolites and natural product therapeutics. Most NRPS biosynthetic pathways include an NRPS and additional proteins that introduce chemical modifications before, during or after assembly-line synthesis. The bacillamide biosynthetic pathway is a common, three-protein system, with a decarboxylase that prepares an NRPS substrate, an NRPS, and an oxidase. Here, the pathway is reconstituted in vitro. The oxidase is shown to perform dehydrogenation of the thiazoline in the peptide intermediate while it is covalently attached to the NRPS, as the penultimate step in bacillamide D synthesis. Structural analysis of the oxidase reveals a dimeric, two-lobed architecture with a remnant RiPP recognition element and a dramatic wrapping loop. The oxidase forms a stable complex with the NRPS and dimerizes it. We visualized co-complexes of the oxidase bound to the elongation module of the NRPS using X-ray crystallography and cryo-EM. The three active sites (for adenylation, condensation/cyclization, and oxidation) form an elegant arc to facilitate substrate delivery. The structures enabled a proof-of-principle bioengineering experiment in which the BmdC oxidase domain is embedded into the NRPS. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23588.map.gz emd_23588.map.gz | 398.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23588-v30.xml emd-23588-v30.xml emd-23588.xml emd-23588.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23588_fsc.xml emd_23588_fsc.xml | 16.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_23588.png emd_23588.png | 147.5 KB | ||
| Masks |  emd_23588_msk_1.map emd_23588_msk_1.map | 421.9 MB |  Mask map Mask map | |
| Others |  emd_23588_half_map_1.map.gz emd_23588_half_map_1.map.gz emd_23588_half_map_2.map.gz emd_23588_half_map_2.map.gz | 391.5 MB 391.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23588 http://ftp.pdbj.org/pub/emdb/structures/EMD-23588 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23588 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23588 | HTTPS FTP |
-Validation report
| Summary document |  emd_23588_validation.pdf.gz emd_23588_validation.pdf.gz | 560.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23588_full_validation.pdf.gz emd_23588_full_validation.pdf.gz | 560.3 KB | Display | |
| Data in XML |  emd_23588_validation.xml.gz emd_23588_validation.xml.gz | 25.1 KB | Display | |
| Data in CIF |  emd_23588_validation.cif.gz emd_23588_validation.cif.gz | 32.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23588 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23588 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23588 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23588 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_23588.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23588.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM Map of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.14 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23588_msk_1.map emd_23588_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
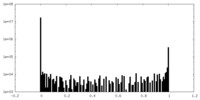
| Density Histograms |
-Half map: Half Map B of the elongation module of...
| File | emd_23588_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map B of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC | ||||||||||||
| Projections & Slices |
| ||||||||||||
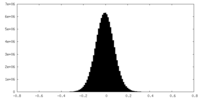
| Density Histograms |
-Half map: Half Map A of the elongation module of...
| File | emd_23588_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map A of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC | ||||||||||||
| Projections & Slices |
| ||||||||||||
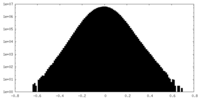
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM map of the elongation module of the bacillamide NRPS, Bmd...
| Entire | Name: Cryo-EM map of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM map of the elongation module of the bacillamide NRPS, Bmd...
| Supramolecule | Name: Cryo-EM map of the elongation module of the bacillamide NRPS, BmdB, complexed with the oxidase BmdC type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Thermoactinomyces vulgaris (bacteria) Thermoactinomyces vulgaris (bacteria) |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 109.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)