[English] 日本語
 Yorodumi
Yorodumi- EMDB-21409: Mycobacterium tuberculosis RNAP S456L mutant open promoter complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21409 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mycobacterium tuberculosis RNAP S456L mutant open promoter complex | |||||||||
 Map data Map data | Mycobacterium tuberculosis RNAP S456L mutant open promoter complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | initiation / transcription bubble / open promoter complex / TRANSCRIPTION / TRANSFERASE-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to water / bacterial-type RNA polymerase holo enzyme binding / Antimicrobial action and antimicrobial resistance in Mtb / sigma factor antagonist complex / rRNA transcription / sigma factor activity / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / peptidoglycan-based cell wall / DNA-directed RNA polymerase complex ...response to water / bacterial-type RNA polymerase holo enzyme binding / Antimicrobial action and antimicrobial resistance in Mtb / sigma factor antagonist complex / rRNA transcription / sigma factor activity / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / peptidoglycan-based cell wall / DNA-directed RNA polymerase complex / DNA-templated transcription initiation / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / nucleic acid binding / protein dimerization activity / transcription cis-regulatory region binding / response to antibiotic / regulation of DNA-templated transcription / DNA-templated transcription / positive regulation of DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.59 Å | |||||||||
 Authors Authors | Lilic M / Boyaci H | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2020 Journal: Proc Natl Acad Sci U S A / Year: 2020Title: The antibiotic sorangicin A inhibits promoter DNA unwinding in a rifampicin-resistant RNA polymerase. Authors: Mirjana Lilic / James Chen / Hande Boyaci / Nathaniel Braffman / Elizabeth A Hubin / Jennifer Herrmann / Rolf Müller / Rachel Mooney / Robert Landick / Seth A Darst / Elizabeth A Campbell /   Abstract: Rifampicin (Rif) is a first-line therapeutic used to treat the infectious disease tuberculosis (TB), which is caused by the pathogen (). The emergence of Rif-resistant (Rif) presents a need for new ...Rifampicin (Rif) is a first-line therapeutic used to treat the infectious disease tuberculosis (TB), which is caused by the pathogen (). The emergence of Rif-resistant (Rif) presents a need for new antibiotics. Rif targets the enzyme RNA polymerase (RNAP). Sorangicin A (Sor) is an unrelated inhibitor that binds in the Rif-binding pocket of RNAP. Sor inhibits a subset of Rif RNAPs, including the most prevalent clinical Rif RNAP substitution found in infected patients (S456>L of the β subunit). Here, we present structural and biochemical data demonstrating that Sor inhibits the wild-type RNAP by a similar mechanism as Rif: by preventing the translocation of very short RNAs. By contrast, Sor inhibits the Rif S456L enzyme at an earlier step, preventing the transition of a partially unwound promoter DNA intermediate to the fully opened DNA and blocking the template-strand DNA from reaching the active site in the RNAP catalytic center. By defining template-strand blocking as a mechanism for inhibition, we provide a mechanistic drug target in RNAP. Our finding that Sor inhibits the wild-type and mutant RNAPs through different mechanisms prompts future considerations for designing antibiotics against resistant targets. Also, we show that Sor has a better pharmacokinetic profile than Rif, making it a suitable starting molecule to design drugs to be used for the treatment of TB patients with comorbidities who require multiple medications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21409.map.gz emd_21409.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21409-v30.xml emd-21409-v30.xml emd-21409.xml emd-21409.xml | 26.9 KB 26.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21409.png emd_21409.png | 165.6 KB | ||
| Filedesc metadata |  emd-21409.cif.gz emd-21409.cif.gz | 9.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21409 http://ftp.pdbj.org/pub/emdb/structures/EMD-21409 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21409 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21409 | HTTPS FTP |
-Related structure data
| Related structure data |  6vw0MC  6vvsC  6vvtC  6vvvC  6vvxC  6vvyC  6vvzC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21409.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21409.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium tuberculosis RNAP S456L mutant open promoter complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
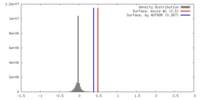
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Mycobacterium tuberculosis RNAP S456L mutant open promoter complex
+Supramolecule #1: Mycobacterium tuberculosis RNAP S456L mutant open promoter complex
+Macromolecule #1: DNA-directed RNA polymerase subunit alpha
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase subunit beta'
+Macromolecule #4: DNA-directed RNA polymerase subunit omega
+Macromolecule #5: RNA polymerase sigma factor SigA
+Macromolecule #6: RNA polymerase-binding protein RbpA
+Macromolecule #7: RNA polymerase-binding transcription factor CarD
+Macromolecule #8: DNA (65-MER)
+Macromolecule #9: DNA (65-MER)
+Macromolecule #10: ZINC ION
+Macromolecule #11: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 71.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)