+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20624 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Empty AAV3B | |||||||||
 Map data Map data | Empty AAV3B (No genome) | |||||||||
 Sample Sample | Parvoviridae != Adeno-associated virus 3B Parvoviridae
| |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Adeno-associated virus 3B Adeno-associated virus 3B | |||||||||
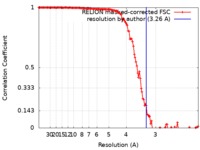
| Method | single particle reconstruction / cryo EM / Resolution: 3.26 Å | |||||||||
 Authors Authors | Subramanian S / Hafenstein S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Hum Gene Ther / Year: 2019 Journal: Hum Gene Ther / Year: 2019Title: Filling Adeno-Associated Virus Capsids: Estimating Success by Cryo-Electron Microscopy. Authors: Suriyasri Subramanian / Anna C Maurer / Carol M Bator / Alexander M Makhov / James F Conway / Kevin B Turner / James H Marden / Luk H Vandenberghe / Susan L Hafenstein /  Abstract: Adeno-associated viruses (AAVs) have been employed successfully as gene therapy vectors in treating various genetic diseases for almost two decades. However, transgene packaging is usually imperfect, ...Adeno-associated viruses (AAVs) have been employed successfully as gene therapy vectors in treating various genetic diseases for almost two decades. However, transgene packaging is usually imperfect, and developing a rapid and accurate method for measuring the proportion of DNA encapsidation is an important step for improving the downstream process of large scale vector production. In this study, we used two-dimensional class averages and three-dimensional classes, intermediate outputs in the single particle cryo-electron microscopy (cryo-EM) image reconstruction pipeline, to determine the proportion of DNA-packaged and empty capsid populations. Two different preparations of AAV3 were analyzed to estimate the minimum number of particles required to be sampled by cryo-EM in order for robust calculation of the proportion of the full versus empty capsids in any given sample. Cost analysis applied to the minimum amount of data required for a valid ratio suggests that cryo-EM is an effective approach to analyze vector preparations. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20624.map.gz emd_20624.map.gz | 181 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20624-v30.xml emd-20624-v30.xml emd-20624.xml emd-20624.xml | 12.5 KB 12.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20624_fsc.xml emd_20624_fsc.xml | 13.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_20624.png emd_20624.png | 232.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20624 http://ftp.pdbj.org/pub/emdb/structures/EMD-20624 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20624 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20624 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20624.map.gz / Format: CCP4 / Size: 193.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20624.map.gz / Format: CCP4 / Size: 193.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Empty AAV3B (No genome) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.136 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Parvoviridae
| Entire | Name:  Parvoviridae (virus) Parvoviridae (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Adeno-associated virus 3B
| Supramolecule | Name: Adeno-associated virus 3B / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 68742 / Sci species name: Adeno-associated virus 3B / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  |
| Host system | Organism:  Homo sapiens (human) / Recombinant cell: HEK Homo sapiens (human) / Recombinant cell: HEK |
| Virus shell | Shell ID: 1 / Diameter: 250.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: AAV3B
| Macromolecule | Name: AAV3B / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAADGYLPDW LEDNLSEGIR EWWALKPGVP QPKANQQHQD NRRGLVLPGY KYLGPGNGLD KGEPVNEAD AAALEHDKAY DQQLKAGDNP YLKYNHADAE FQERLQEDTS FGGNLGRAVF Q AKKRILEP LGLVEEAAKT APGKKRPVDQ SPQEPDSSSG VGKSGKQPAR ...String: MAADGYLPDW LEDNLSEGIR EWWALKPGVP QPKANQQHQD NRRGLVLPGY KYLGPGNGLD KGEPVNEAD AAALEHDKAY DQQLKAGDNP YLKYNHADAE FQERLQEDTS FGGNLGRAVF Q AKKRILEP LGLVEEAAKT APGKKRPVDQ SPQEPDSSSG VGKSGKQPAR KRLNFGQTGD SE SVPDPQP LGEPPAAPTS LGSNTMASGG GAPMADNNEG ADGVGNSSGN WHCDSQWLGD RVI TTSTRT WALPTYNNHL YKQISSQSGA SNDNHYFGYS TPWGYFDFNR FHCHFSPRDW QRLI NNNWG FRPKKLSFKL FNIQVKEVTQ NDGTTTIANN LTSTVQVFTD SEYQLPYVLG SAHQG CLPP FPADVFMVPQ YGYLTLNNGS QAVGRSSFYC LEYFPSQMLR TGNNFQFSYT FEDVPF HSS YAHSQSLDRL MNPLIDQYLY YLNRTQGTTS GTTNQSRLLF SQAGPQSMSL QARNWLP GP CYRQQRLSKT ANDNNNSNFP WTAASKYHLN GRDSLVNPGP AMASHKDDEE KFFPMHGN L IFGKEGTTAS NAELDNVMIT DEEEIRTTNP VATEQYGTVA NNLQSSNTAP TTRTVNDQG ALPGMVWQDR DVYLQGPIWA KIPHTDGHFH PSPLMGGFGL KHPPPQIMIK NTPVPANPPT TFSPAKFAS FITQYSTGQV SVEIEWELQK ENSKRWNPEI QYTSNYNKSV NVDFTVDTNG V YSEPRPIG TRYLTRNL |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
Details: Phosphate buffered saline pH 7.4 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Details: unspecified | |||||||||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number real images: 2082 / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)