[English] 日本語
 Yorodumi
Yorodumi- EMDB-20354: cryoEM structure of the substrate-bound human CTP synthase 2 filament -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20354 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryoEM structure of the substrate-bound human CTP synthase 2 filament | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Enzyme / filament / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationCTP synthase (glutamine hydrolysing) / CTP synthase activity / pyrimidine nucleotide metabolic process / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / Interconversion of nucleotide di- and triphosphates / CTP biosynthetic process / mitochondrion / ATP binding ...CTP synthase (glutamine hydrolysing) / CTP synthase activity / pyrimidine nucleotide metabolic process / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / Interconversion of nucleotide di- and triphosphates / CTP biosynthetic process / mitochondrion / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Lynch EM / Kollman JM | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2020 Journal: Nat Struct Mol Biol / Year: 2020Title: Coupled structural transitions enable highly cooperative regulation of human CTPS2 filaments. Authors: Eric M Lynch / Justin M Kollman /  Abstract: Many enzymes assemble into defined oligomers, providing a mechanism for cooperatively regulating activity. Recent studies have described a mode of regulation in which enzyme activity is modulated by ...Many enzymes assemble into defined oligomers, providing a mechanism for cooperatively regulating activity. Recent studies have described a mode of regulation in which enzyme activity is modulated by polymerization into large-scale filaments. Here we describe an ultrasensitive form of polymerization-based regulation employed by human CTP synthase 2 (CTPS2). Cryo-EM structures reveal that CTPS2 filaments dynamically switch between active and inactive forms in response to changes in substrate and product levels. Linking the conformational state of many CTPS2 subunits in a filament results in highly cooperative regulation, greatly exceeding the limits of cooperativity for the CTPS2 tetramer alone. The structures reveal a link between conformation and control of ammonia channeling between the enzyme's active sites, and explain differences in regulation of human CTPS isoforms. This filament-based mechanism of enhanced cooperativity demonstrates how the widespread phenomenon of enzyme polymerization can be adapted to achieve different regulatory outcomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20354.map.gz emd_20354.map.gz | 95.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20354-v30.xml emd-20354-v30.xml emd-20354.xml emd-20354.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20354.png emd_20354.png | 245.4 KB | ||
| Filedesc metadata |  emd-20354.cif.gz emd-20354.cif.gz | 5.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20354 http://ftp.pdbj.org/pub/emdb/structures/EMD-20354 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20354 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20354 | HTTPS FTP |
-Related structure data
| Related structure data |  6pk4MC  6pk7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20354.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20354.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
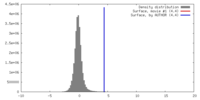
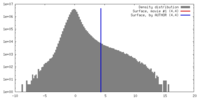
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : substrate-bound CTP synthase 2 filament
| Entire | Name: substrate-bound CTP synthase 2 filament |
|---|---|
| Components |
|
-Supramolecule #1: substrate-bound CTP synthase 2 filament
| Supramolecule | Name: substrate-bound CTP synthase 2 filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: CTP synthase 2
| Macromolecule | Name: CTP synthase 2 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: CTP synthase (glutamine hydrolysing) |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 65.75943 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYILVTGGV ISGIGKGIIA SSIGTILKSC GLRVTAIKID PYINIDAGTF SPYEHGEVFV LNDGGEVDLD LGNYERFLDI NLYKDNNIT TGKIYQHVIN KERRGDYLGK TVQVVPHITD AVQEWVMNQA KVPVDGNKEE PQICVIELGG TIGDIEGMPF V EAFRQFQF ...String: MKYILVTGGV ISGIGKGIIA SSIGTILKSC GLRVTAIKID PYINIDAGTF SPYEHGEVFV LNDGGEVDLD LGNYERFLDI NLYKDNNIT TGKIYQHVIN KERRGDYLGK TVQVVPHITD AVQEWVMNQA KVPVDGNKEE PQICVIELGG TIGDIEGMPF V EAFRQFQF KAKRENFCNI HVSLVPQLSA TGEQKTKPTQ NSVRALRGLG LSPDLIVCRS STPIEMAVKE KISMFCHVNP EQ VICIHDV SSTYRVPVLL EEQSIVKYFK ERLHLPIGDS ASNLLFKWRN MADRYERLQK ICSIALVGKY TKLRDCYASV FKA LEHSAL AINHKLNLMY IDSIDLEKIT ETEDPVKFHE AWQKLCKADG ILVPGGFGIR GTLGKLQAIS WARTKKIPFL GVCL GMQLA VIEFARNCLN LKDADSTEFR PNAPVPLVID MPEHNPGNLG GTMRLGIRRT VFKTENSILR KLYGDVPFIE ERHRH RFEV NPNLIKQFEQ NDLSFVGQDV DGDRMEIIEL ANHPYFVGVQ FHPEFSSRPM KPSPPYLGLL LAATGNLNAY LQQGCK LSS SDRYSDASDD SFSEPRIAEL EIS UniProtKB: CTP synthase 2 |
-Macromolecule #2: URIDINE 5'-TRIPHOSPHATE
| Macromolecule | Name: URIDINE 5'-TRIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 4 / Formula: UTP |
|---|---|
| Molecular weight | Theoretical: 484.141 Da |
| Chemical component information |  ChemComp-UTP: |
-Macromolecule #3: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 90.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Applied symmetry - Point group: D2 (2x2 fold dihedral) / Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cisTEM / Number images used: 53964 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cisTEM |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)