[English] 日本語
 Yorodumi
Yorodumi- EMDB-20055: Cryo-EM structure of Her2 extracellular domain-Trastuzumab Fab-Pe... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20055 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Her2 extracellular domain-Trastuzumab Fab-Pertuzumab Fab complex | |||||||||
 Map data Map data | Sharpened map with phenix.auto_sharpen | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Her2 extracellular domain / Trastuzumab / Pertuzumab / transferase-immune system complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of immature T cell proliferation in thymus / IgD immunoglobulin complex / ERBB3:ERBB2 complex / ERBB2-ERBB4 signaling pathway / immature T cell proliferation in thymus / GRB7 events in ERBB2 signaling / IgA immunoglobulin complex / IgM immunoglobulin complex / IgE immunoglobulin complex / RNA polymerase I core binding ...negative regulation of immature T cell proliferation in thymus / IgD immunoglobulin complex / ERBB3:ERBB2 complex / ERBB2-ERBB4 signaling pathway / immature T cell proliferation in thymus / GRB7 events in ERBB2 signaling / IgA immunoglobulin complex / IgM immunoglobulin complex / IgE immunoglobulin complex / RNA polymerase I core binding / semaphorin receptor complex / Developmental Lineage of Mammary Stem Cells / CD22 mediated BCR regulation / ErbB-3 class receptor binding / motor neuron axon guidance / Sema4D induced cell migration and growth-cone collapse / Fc epsilon receptor (FCERI) signaling / regulation of microtubule-based process / Classical antibody-mediated complement activation / Initial triggering of complement / IgG immunoglobulin complex / PLCG1 events in ERBB2 signaling / enzyme-linked receptor protein signaling pathway / ERBB2 Activates PTK6 Signaling / ERBB2-EGFR signaling pathway / ERBB2-ERBB3 signaling pathway / Drug-mediated inhibition of ERBB2 signaling / Resistance of ERBB2 KD mutants to trastuzumab / Resistance of ERBB2 KD mutants to sapitinib / Resistance of ERBB2 KD mutants to tesevatinib / Resistance of ERBB2 KD mutants to neratinib / Resistance of ERBB2 KD mutants to osimertinib / Resistance of ERBB2 KD mutants to afatinib / Resistance of ERBB2 KD mutants to AEE788 / Resistance of ERBB2 KD mutants to lapatinib / Drug resistance in ERBB2 TMD/JMD mutants / neurotransmitter receptor localization to postsynaptic specialization membrane / positive regulation of MAP kinase activity / positive regulation of Rho protein signal transduction / neuromuscular junction development / positive regulation of transcription by RNA polymerase I / immunoglobulin mediated immune response / ERBB2 Regulates Cell Motility / Developmental Lineage of Mammary Gland Myoepithelial Cells / oligodendrocyte differentiation / semaphorin-plexin signaling pathway / FCGR activation / PI3K events in ERBB2 signaling / Role of LAT2/NTAL/LAB on calcium mobilization / regulation of angiogenesis / Role of phospholipids in phagocytosis / positive regulation of protein targeting to membrane / immunoglobulin complex / Scavenging of heme from plasma / regulation of ERK1 and ERK2 cascade / Schwann cell development / antigen binding / coreceptor activity / peptidyl-tyrosine phosphorylation / Signaling by ERBB2 / TFAP2 (AP-2) family regulates transcription of growth factors and their receptors / myelination / transmembrane receptor protein tyrosine kinase activity / FCERI mediated Ca+2 mobilization / GRB2 events in ERBB2 signaling / FCGR3A-mediated IL10 synthesis / positive regulation of cell adhesion / SHC1 events in ERBB2 signaling / Antigen activates B Cell Receptor (BCR) leading to generation of second messengers / Regulation of Complement cascade / cell surface receptor protein tyrosine kinase signaling pathway / cellular response to epidermal growth factor stimulus / Constitutive Signaling by Overexpressed ERBB2 / basal plasma membrane / Downregulation of ERBB2:ERBB3 signaling / positive regulation of epithelial cell proliferation / Cell surface interactions at the vascular wall / B cell receptor signaling pathway / positive regulation of translation / FCGR3A-mediated phagocytosis / FCERI mediated MAPK activation / neuromuscular junction / wound healing / phosphatidylinositol 3-kinase/protein kinase B signal transduction / Signaling by ERBB2 TMD/JMD mutants / receptor protein-tyrosine kinase / Signaling by ERBB2 ECD mutants / Signaling by ERBB2 KD Mutants / Regulation of actin dynamics for phagocytic cup formation / receptor tyrosine kinase binding / cellular response to growth factor stimulus / Downregulation of ERBB2 signaling / FCERI mediated NF-kB activation / ruffle membrane / epidermal growth factor receptor signaling pathway / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / neuron differentiation / Constitutive Signaling by Aberrant PI3K in Cancer / transmembrane signaling receptor activity / PIP3 activates AKT signaling Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.36 Å | |||||||||
 Authors Authors | Hao Y / Yu X | |||||||||
 Citation Citation |  Journal: PLoS One / Year: 2019 Journal: PLoS One / Year: 2019Title: Cryo-EM Structure of HER2-trastuzumab-pertuzumab complex. Authors: Yue Hao / Xinchao Yu / Yonghong Bai / Helen J McBride / Xin Huang /  Abstract: Trastuzumab and pertuzumab are monoclonal antibodies that bind to distinct subdomains of the extracellular domain of human epidermal growth factor receptor 2 (HER2). Adding these monoclonal ...Trastuzumab and pertuzumab are monoclonal antibodies that bind to distinct subdomains of the extracellular domain of human epidermal growth factor receptor 2 (HER2). Adding these monoclonal antibodies to the treatment regimen of HER2-positive breast cancer has changed the paradigm for treatment in that form of cancer. Synergistic activity has been observed with the combination of these two antibodies leading to hypotheses regarding the mechanism(s) and to the development of bispecific antibodies to maximize the clinical effect further. Although the individual crystal structures of HER2-trastuzumab and HER2-pertuzumab revealed the distinct binding sites and provided the structural basis for their anti-tumor activities, detailed structural information on the HER2-trastuzumab-pertuzumab complex has been elusive. Here we present the cryo-EM structure of HER2-trastuzumab-pertuzumab at 4.36 Å resolution. Comparison with the binary complexes reveals no cooperative interaction between trastuzumab and pertuzumab, and provides key insights into the design of novel, high-avidity bispecific molecules with potentially greater clinical efficacy. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20055.map.gz emd_20055.map.gz | 31.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20055-v30.xml emd-20055-v30.xml emd-20055.xml emd-20055.xml | 18.4 KB 18.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20055.png emd_20055.png | 145.4 KB | ||
| Filedesc metadata |  emd-20055.cif.gz emd-20055.cif.gz | 6.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20055 http://ftp.pdbj.org/pub/emdb/structures/EMD-20055 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20055 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20055 | HTTPS FTP |
-Related structure data
| Related structure data |  6ogeMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20055.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20055.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map with phenix.auto_sharpen | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
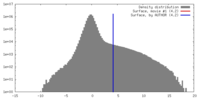
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Her2 extracellular domain-Trastuzumab Fab-Pertuzumab Fab complex
+Supramolecule #1: Her2 extracellular domain-Trastuzumab Fab-Pertuzumab Fab complex
+Supramolecule #2: Human HER2 extracellular domain
+Supramolecule #3: Pertuzumab Fab
+Supramolecule #4: Trastuzumab Fab
+Macromolecule #1: Receptor tyrosine-protein kinase erbB-2
+Macromolecule #2: Pertuzumab FAB LIGHT CHAIN
+Macromolecule #3: Pertuzumab FAB HEAVY CHAIN
+Macromolecule #4: Trastuzumab FAB LIGHT CHAIN
+Macromolecule #5: Trastuzumab FAB HEAVY CHAIN
+Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.4 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Mesh: 300 | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average exposure time: 6.0 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -1.5 µm / Nominal defocus min: -3.5 µm / Nominal magnification: 130000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6oge: |
 Movie
Movie Controller
Controller






































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)