[English] 日本語
 Yorodumi
Yorodumi- EMDB-18864: Central glycolytic genes regulator (CggR) bound to DNA operator -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Central glycolytic genes regulator (CggR) bound to DNA operator | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacillus subtilis / transcription repressor / glucose catabolism / SorC family / DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of DNA-templated transcription initiation / cis-regulatory region sequence-specific DNA binding / carbohydrate binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
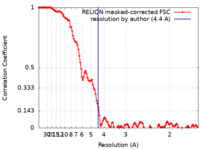
| Method | single particle reconstruction / cryo EM / Resolution: 4.4 Å | |||||||||
 Authors Authors | Skerlova J / Soltysova M / Rezacova P / Skubnik K | |||||||||
| Funding support |  Czech Republic, European Union, 2 items Czech Republic, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2024 Journal: Nucleic Acids Res / Year: 2024Title: Structural characterization of two prototypical repressors of SorC family reveals tetrameric assemblies on DNA and mechanism of function. Authors: Markéta Šoltysová / Jana Škerlová / Petr Pachl / Karel Škubník / Milan Fábry / Irena Sieglová / Martina Farolfi / Irina Grishkovskaya / Michal Babiak / Jiří Nováček / Libor ...Authors: Markéta Šoltysová / Jana Škerlová / Petr Pachl / Karel Škubník / Milan Fábry / Irena Sieglová / Martina Farolfi / Irina Grishkovskaya / Michal Babiak / Jiří Nováček / Libor Krásný / Pavlína Řezáčová /  Abstract: The SorC family of transcriptional regulators plays a crucial role in controlling the carbohydrate metabolism and quorum sensing. We employed an integrative approach combining X-ray crystallography ...The SorC family of transcriptional regulators plays a crucial role in controlling the carbohydrate metabolism and quorum sensing. We employed an integrative approach combining X-ray crystallography and cryo-electron microscopy to investigate architecture and functional mechanism of two prototypical representatives of two sub-classes of the SorC family: DeoR and CggR from Bacillus subtilis. Despite possessing distinct DNA-binding domains, both proteins form similar tetrameric assemblies when bound to their respective DNA operators. Structural analysis elucidates the process by which the CggR-regulated gapA operon is derepressed through the action of two effectors: fructose-1,6-bisphosphate and newly confirmed dihydroxyacetone phosphate. Our findings provide the first comprehensive understanding of the DNA binding mechanism of the SorC-family proteins, shedding new light on their functional characteristics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18864.map.gz emd_18864.map.gz | 156.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18864-v30.xml emd-18864-v30.xml emd-18864.xml emd-18864.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18864_fsc.xml emd_18864_fsc.xml | 12.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_18864.png emd_18864.png | 78.8 KB | ||
| Masks |  emd_18864_msk_1.map emd_18864_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18864.cif.gz emd-18864.cif.gz | 7 KB | ||
| Others |  emd_18864_half_map_1.map.gz emd_18864_half_map_1.map.gz emd_18864_half_map_2.map.gz emd_18864_half_map_2.map.gz | 154.6 MB 154.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18864 http://ftp.pdbj.org/pub/emdb/structures/EMD-18864 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18864 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18864 | HTTPS FTP |
-Related structure data
| Related structure data |  8r3gMC  8r7yC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18864.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18864.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||
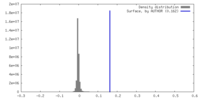
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18864_msk_1.map emd_18864_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18864_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
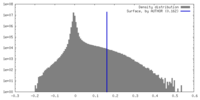
| Density Histograms |
-Half map: #1
| File | emd_18864_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
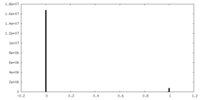
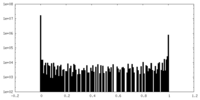
| Density Histograms |
- Sample components
Sample components
-Entire : A complex of Central glycolytic genes regulator (CggR) from Bacil...
| Entire | Name: A complex of Central glycolytic genes regulator (CggR) from Bacillus subtilis and the DNA operator (OLR). |
|---|---|
| Components |
|
-Supramolecule #1: A complex of Central glycolytic genes regulator (CggR) from Bacil...
| Supramolecule | Name: A complex of Central glycolytic genes regulator (CggR) from Bacillus subtilis and the DNA operator (OLR). type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: A complex of full-length CggR with its 45bp DNA operator. |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Central glycolytic genes regulator
| Macromolecule | Name: Central glycolytic genes regulator / type: protein_or_peptide / ID: 1 Details: N-terminal amino-acid sequence GIDPFT remains a part of the protein upon TEV cleavage as a cloning artefact. Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.623496 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GIDPFT(MSE)NQL IQAQKKLLPD LLLV(MSE)QKRFE ILQYIRLTEP IGRRSLSASL GISERVLRGE VQFLKEQNLV DI KTNG(MSE)TL TEEGYELLSV LEDT(MSE)KDVLG LTLLEKTLKE RLNLKDAIIV SGDSDQSPWV KKE(MSE)GRAAVA C (MSE)KKRFSGK ...String: GIDPFT(MSE)NQL IQAQKKLLPD LLLV(MSE)QKRFE ILQYIRLTEP IGRRSLSASL GISERVLRGE VQFLKEQNLV DI KTNG(MSE)TL TEEGYELLSV LEDT(MSE)KDVLG LTLLEKTLKE RLNLKDAIIV SGDSDQSPWV KKE(MSE)GRAAVA C (MSE)KKRFSGK NIVAVTGGTT IEAVAE(MSE)(MSE)TP DSKNRELLFV PARGGLGEDV KNQANTICAH (MSE)AEKAS GTY RLLFVPGQLS QGAYSSIIEE PSVKEVLNTI KSAS(MSE)LVHGI GEAKT(MSE)AQRR NTPLEDLKKI DDNDAVTEA FGYYFNADGE VVHKVHSVG(MSE) QLDDIDAIPD IIAVAGGSSK AEAIEAYFKK PRNTVLVTDE GAAKKLLRDE UniProtKB: Central glycolytic genes regulator |
-Macromolecule #2: operator DNA
| Macromolecule | Name: operator DNA / type: dna / ID: 2 Details: Only 41bp were modelled into the electron density map (two base pairs at both ends are missing). Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.78087 KDa |
| Sequence | String: (DT)(DT)(DG)(DC)(DT)(DG)(DG)(DA)(DC)(DA) (DT)(DT)(DA)(DT)(DA)(DT)(DG)(DT)(DC)(DC) (DC)(DG)(DC)(DT)(DA)(DT)(DG)(DA)(DC) (DA)(DA)(DA)(DA)(DA)(DA)(DC)(DG)(DT)(DC) (DC) (DC)(DG)(DT)(DC)(DA) |
-Macromolecule #3: operator DNA
| Macromolecule | Name: operator DNA / type: dna / ID: 3 Details: Only 41bp were modelled into the electron density map (two base pairs at both sides are missing). Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.931952 KDa |
| Sequence | String: (DT)(DG)(DA)(DC)(DG)(DG)(DG)(DA)(DC)(DG) (DT)(DT)(DT)(DT)(DT)(DT)(DG)(DT)(DC)(DA) (DT)(DA)(DG)(DC)(DG)(DG)(DG)(DA)(DC) (DA)(DT)(DA)(DT)(DA)(DA)(DT)(DG)(DT)(DC) (DC) (DA)(DG)(DC)(DA)(DA) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.86 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 20 mM Tris-HCl, pH 7.5, 100 mM NaCl and 0.02% (v/v) beta-mercaptoethanol | ||||||||||||
| Grid | Model: C-flat / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 30.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Details | Rigid body fitting of starting PDB models in Phenix and Coot was combined with manual rebuilding of certain model regions in Coot. | ||||||
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 24.32 | ||||||
| Output model |  PDB-8r3g: |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)