[English] 日本語
 Yorodumi
Yorodumi- EMDB-17841: Cryo-EM structure of Sodium proton exchanger NhaA with bound card... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Sodium proton exchanger NhaA with bound cardiolipin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Sodium proton exchanger / Cardiolipin / NhaA / TRANSPORT PROTEIN | |||||||||
| Function / homology | Na+/H+ antiporter NhaA / Na+/H+ antiporter domain superfamily / Na+/H+ antiporter 1 / sodium:proton antiporter activity / regulation of pH / plasma membrane / Na(+)/H(+) antiporter NhaA Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
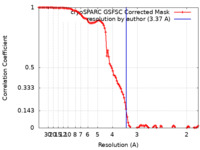
| Method | single particle reconstruction / cryo EM / Resolution: 3.37 Å | |||||||||
 Authors Authors | Gulati A / Meier P / Kokane S / Drew D | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2025 Journal: Nat Commun / Year: 2025Title: PIP-mediated oligomerization of the endosomal sodium/proton exchanger NHE9. Authors: Surabhi Kokane / Ashutosh Gulati / Pascal F Meier / Rei Matsuoka / Tanadet Pipatpolkai / Giuseppe Albano / Tin Manh Ho / Lucie Delemotte / Daniel Fuster / David Drew /   Abstract: The strict exchange of Na for H ions across cell membranes is a reaction carried out in almost every cell. Na/H exchangers that perform this task are physiological homodimers, and whilst the ion ...The strict exchange of Na for H ions across cell membranes is a reaction carried out in almost every cell. Na/H exchangers that perform this task are physiological homodimers, and whilst the ion transporting domain is highly conserved, their dimerization differs. The Na/H exchanger NhaA from Escherichia coli has a weak dimerization interface mediated by a β-hairpin domain and with dimer retention dependent on cardiolipin. Similarly, organellar Na/H exchangers NHE6, NHE7 and NHE9 also contain β-hairpin domains and recent analysis of Equus caballus NHE9 indicated PIP lipids could bind at the dimer interface. However, structural validation of the predicted lipid-mediated oligomerization has been lacking. Here, we report cryo-EM structures of E. coli NhaA and E. caballus NHE9 in complex with cardiolipin and phosphatidylinositol-3,5-bisphosphate PI(3,5)P lipids binding at their respective dimer interfaces. We further show how the endosomal specific PI(3,5)P lipid stabilizes the NHE9 homodimer and enhances transport activity. Indeed, we show that NHE9 is active in endosomes, but not at the plasma membrane where the PI(3,5)P lipid is absent. Thus, specific lipids can regulate Na/H exchange activity by stabilizing dimerization in response to either cell specific cues or upon trafficking to their correct membrane location. | |||||||||
| History |
|
- Structure visualization
Structure visualization
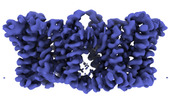
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17841.map.gz emd_17841.map.gz | 106.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17841-v30.xml emd-17841-v30.xml emd-17841.xml emd-17841.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17841_fsc.xml emd_17841_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_17841.png emd_17841.png | 104.8 KB | ||
| Masks |  emd_17841_msk_1.map emd_17841_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17841.cif.gz emd-17841.cif.gz | 6 KB | ||
| Others |  emd_17841_half_map_1.map.gz emd_17841_half_map_1.map.gz emd_17841_half_map_2.map.gz emd_17841_half_map_2.map.gz | 115.6 MB 115.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17841 http://ftp.pdbj.org/pub/emdb/structures/EMD-17841 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17841 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17841 | HTTPS FTP |
-Related structure data
| Related structure data |  8ps0MC  8pvrC  8pxbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17841.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17841.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9137 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17841_msk_1.map emd_17841_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17841_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17841_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : NhaA dimer
| Entire | Name: NhaA dimer |
|---|---|
| Components |
|
-Supramolecule #1: NhaA dimer
| Supramolecule | Name: NhaA dimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Na(+)/H(+) antiporter NhaA
| Macromolecule | Name: Na(+)/H(+) antiporter NhaA / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.304422 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKHLHRFFSS DASGGIILII AAILAMIMAN SGATSGWYHD FLETPVQLRV GSLEINKNML LWINDALMAV FFLLVGLEVK RELMQGSLA SLRQAAFPVI AAIGGMIVPT LLYLAFNYAD PITREGWAIP AATDIAFALG VLALLGSRVP LALKIFLMAL A IIDDLGAI ...String: MKHLHRFFSS DASGGIILII AAILAMIMAN SGATSGWYHD FLETPVQLRV GSLEINKNML LWINDALMAV FFLLVGLEVK RELMQGSLA SLRQAAFPVI AAIGGMIVPT LLYLAFNYAD PITREGWAIP AATDIAFALG VLALLGSRVP LALKIFLMAL A IIDDLGAI IIIALFYTND LSMASLGVAA VAIAVLAVLN LCGARRTGVY ILVGVVLWTA VLKSGVHATL AGVIVGFFIP LK EKHGRSP AKRLEHVLHP WVAYLILPLF AFANAGVSLG GVTLDGLTSI LPLGIIAGML IGKPLGISLF CWLALRLKLA HLP EGTTYQ QIMVVGILCG IGFTMSIFIA SLAFGSVDPE LINWAKLGIL VGSISSAVIG YSWLRVRLRP SVGSENLYFQ UniProtKB: Na(+)/H(+) antiporter NhaA |
-Macromolecule #2: CARDIOLIPIN
| Macromolecule | Name: CARDIOLIPIN / type: ligand / ID: 2 / Number of copies: 3 / Formula: CDL |
|---|---|
| Molecular weight | Theoretical: 1.464043 KDa |
| Chemical component information |  ChemComp-CDL: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 68.11 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)