+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1663 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Model of the Vesicular Stomatitis Virus | |||||||||
 Map data Map data | This is one octant of the volume of the top view of the VSV virion trunk. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NSRV / VSV / CryoEM / RNA / helix | |||||||||
| Function / homology |  Function and homology information Function and homology informationRNA replication / helical viral capsid / viral transcription / viral nucleocapsid / host cell cytoplasm / ribonucleoprotein complex / RNA binding Similarity search - Function | |||||||||
| Biological species |  Vesicular stomatitis virus Vesicular stomatitis virus | |||||||||
| Method | helical reconstruction / cryo EM / negative staining / Resolution: 10.6 Å | |||||||||
 Authors Authors | Ge P / Tsao J / Green TJ / Luo M / Zhou ZH | |||||||||
 Citation Citation |  Journal: Science / Year: 2010 Journal: Science / Year: 2010Title: Cryo-EM model of the bullet-shaped vesicular stomatitis virus. Authors: Peng Ge / Jun Tsao / Stan Schein / Todd J Green / Ming Luo / Z Hong Zhou /  Abstract: Vesicular stomatitis virus (VSV) is a bullet-shaped rhabdovirus and a model system of negative-strand RNA viruses. Through direct visualization by means of cryo-electron microscopy, we show that each ...Vesicular stomatitis virus (VSV) is a bullet-shaped rhabdovirus and a model system of negative-strand RNA viruses. Through direct visualization by means of cryo-electron microscopy, we show that each virion contains two nested, left-handed helices: an outer helix of matrix protein M and an inner helix of nucleoprotein N and RNA. M has a hub domain with four contact sites that link to neighboring M and N subunits, providing rigidity by clamping adjacent turns of the nucleocapsid. Side-by-side interactions between neighboring N subunits are critical for the nucleocapsid to form a bullet shape, and structure-based mutagenesis results support this description. Together, our data suggest a mechanism of VSV assembly in which the nucleocapsid spirals from the tip to become the helical trunk, both subsequently framed and rigidified by the M layer. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1663.map.gz emd_1663.map.gz | 72.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1663-v30.xml emd-1663-v30.xml emd-1663.xml emd-1663.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| Images |  Fig1.b.down.left-500.jpg Fig1.b.down.left-500.jpg | 131.8 KB | ||
| Masks |  emd_1663_msk_1.map emd_1663_msk_1.map emd_1663_msk_2.map emd_1663_msk_2.map | 6.7 MB 5.5 MB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1663 http://ftp.pdbj.org/pub/emdb/structures/EMD-1663 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1663 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1663 | HTTPS FTP |
-Related structure data
| Related structure data |  2wyyMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1663.map.gz / Format: CCP4 / Size: 122.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1663.map.gz / Format: CCP4 / Size: 122.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is one octant of the volume of the top view of the VSV virion trunk. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.532 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Segmentation: This is 5 consequetive N's in the nucleocapsid
| Annotation | This is 5 consequetive N's in the nucleocapsid | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_1663_msk_1.map emd_1663_msk_1.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
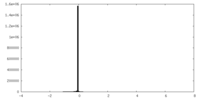
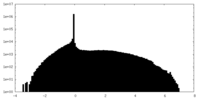
| Density Histograms |
-Segmentation: This is 2N with 5M in closest neighborhood of a central M-hub domain
| Annotation | This is 2N with 5M in closest neighborhood of a central M-hub domain | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_1663_msk_2.map emd_1663_msk_2.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
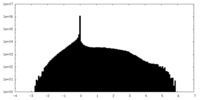
| Density Histograms |
- Sample components
Sample components
-Entire : VSV (Indiana)
| Entire | Name: VSV (Indiana) |
|---|---|
| Components |
|
-Supramolecule #1000: VSV (Indiana)
| Supramolecule | Name: VSV (Indiana) / type: sample / ID: 1000 / Oligomeric state: Mature virion / Number unique components: 1 |
|---|
-Supramolecule #1: Vesicular stomatitis virus
| Supramolecule | Name: Vesicular stomatitis virus / type: virus / ID: 1 / Name.synonym: VSV Details: Horse is the primary host species of the virus but Human is also possible. NCBI-ID: 11276 / Sci species name: Vesicular stomatitis virus / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: VSV |
|---|---|
| Host (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: 100mM NaCl, 10nM Tris, 1mM EDTA |
|---|---|
| Staining | Type: NEGATIVE / Details: Cryo sample |
| Grid | Details: Quantifoil 3.5/1 200 mesh |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 50 % / Chamber temperature: 80 K / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: manual plunger. Vitrification carried out in open room Method: manual blot 1 second before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Temperature | Average: 80 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 250,000 times magnification |
| Date | Jan 5, 2007 |
| Image recording | Category: CCD / Film or detector model: GENERIC TVIPS / Digitization - Sampling interval: 1.532 µm / Number real images: 212 / Average electron dose: 20 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 98000 / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 10.6 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN / Details: Helicity determined and applied by IHRSR |
|---|
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | Protocol: Rigid Body |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross-correlation |
| Output model |  PDB-2wyy: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)