[English] 日本語
 Yorodumi
Yorodumi- EMDB-16172: Cryo-EM structure of the Arabidopsis thaliana I+III2 supercomplex... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Arabidopsis thaliana I+III2 supercomplex (Complete conformation 2 composition) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Plant / Mitochondria / Complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationplastid outer membrane / anther dehiscence / signal peptidase activity / TIM22 mitochondrial import inner membrane insertion complex / vegetative to reproductive phase transition of meristem / chloroplast outer membrane / cold acclimation / mitochondrial processing peptidase / Lyases; Carbon-oxygen lyases; Hydro-lyases / P450-containing electron transport chain ...plastid outer membrane / anther dehiscence / signal peptidase activity / TIM22 mitochondrial import inner membrane insertion complex / vegetative to reproductive phase transition of meristem / chloroplast outer membrane / cold acclimation / mitochondrial processing peptidase / Lyases; Carbon-oxygen lyases; Hydro-lyases / P450-containing electron transport chain / plant-type cell wall / photorespiration / embryo development ending in seed dormancy / protein insertion into mitochondrial inner membrane / NADH dehydrogenase complex / plant-type vacuole / response to abscisic acid / ubiquinone biosynthetic process / vacuole / respiratory chain complex III / cellular respiration / quinol-cytochrome-c reductase / cobalt ion binding / quinol-cytochrome-c reductase activity / response to osmotic stress / regulation of reactive oxygen species metabolic process / plastid / mitochondrial electron transport, ubiquinol to cytochrome c / porin activity / pore complex / protein homotrimerization / NADH:ubiquinone reductase (H+-translocating) / mitochondrial electron transport, NADH to ubiquinone / electron transport coupled proton transport / acyl binding / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / oxidoreductase activity, acting on NAD(P)H / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity / acyl carrier activity / quinone binding / ATP synthesis coupled electron transport / response to salt stress / monoatomic ion transport / proton transmembrane transport / aerobic respiration / chloroplast / carbonate dehydratase activity / respiratory electron transport chain / metalloendopeptidase activity / electron transport chain / mitochondrial intermembrane space / 2 iron, 2 sulfur cluster binding / mitochondrial membrane / NAD binding / fatty acid biosynthetic process / FMN binding / peroxisome / 4 iron, 4 sulfur cluster binding / carbohydrate metabolic process / mitochondrial outer membrane / electron transfer activity / oxidoreductase activity / mitochondrial inner membrane / mitochondrial matrix / copper ion binding / heme binding / protein-containing complex binding / nucleolus / protein homodimerization activity / mitochondrion / proteolysis / extracellular region / zinc ion binding / ATP binding / membrane / metal ion binding / identical protein binding / nucleus / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
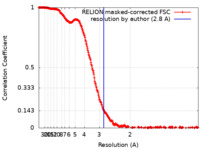
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Klusch N / Kuehlbrandt W | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Plants / Year: 2023 Journal: Nat Plants / Year: 2023Title: Cryo-EM structure of the respiratory I + III supercomplex from Arabidopsis thaliana at 2 Å resolution. Authors: Niklas Klusch / Maximilian Dreimann / Jennifer Senkler / Nils Rugen / Werner Kühlbrandt / Hans-Peter Braun /  Abstract: Protein complexes of the mitochondrial respiratory chain assemble into respiratory supercomplexes. Here we present the high-resolution electron cryo-microscopy structure of the Arabidopsis ...Protein complexes of the mitochondrial respiratory chain assemble into respiratory supercomplexes. Here we present the high-resolution electron cryo-microscopy structure of the Arabidopsis respiratory supercomplex consisting of complex I and a complex III dimer, with a total of 68 protein subunits and numerous bound cofactors. A complex I-ferredoxin, subunit B14.7 and P9, a newly defined subunit of plant complex I, mediate supercomplex formation. The component complexes stabilize one another, enabling new detailed insights into their structure. We describe (1) an interrupted aqueous passage for proton translocation in the membrane arm of complex I; (2) a new coenzyme A within the carbonic anhydrase module of plant complex I defining a second catalytic centre; and (3) the water structure at the proton exit pathway of complex III with a co-purified ubiquinone in the Q site. We propose that the main role of the plant supercomplex is to stabilize its components in the membrane. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16172.map.gz emd_16172.map.gz | 99 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16172-v30.xml emd-16172-v30.xml emd-16172.xml emd-16172.xml | 77.6 KB 77.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16172_fsc.xml emd_16172_fsc.xml | 26.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16172.png emd_16172.png | 186.9 KB | ||
| Filedesc metadata |  emd-16172.cif.gz emd-16172.cif.gz | 16.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16172 http://ftp.pdbj.org/pub/emdb/structures/EMD-16172 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16172 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16172 | HTTPS FTP |
-Related structure data
| Related structure data |  8bq6MC  8bedC  8beeC  8befC  8behC  8belC  8bepC  8bpxC  8bq5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16172.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16172.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.573 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Mitochondrial Arabidopsis thaliana I+III2 supercomplex (Complete ...
+Supramolecule #1: Mitochondrial Arabidopsis thaliana I+III2 supercomplex (Complete ...
+Macromolecule #1: NADH-ubiquinone oxidoreductase chain 3
+Macromolecule #2: NADH dehydrogenase [ubiquinone] iron-sulfur protein 7, mitochondrial
+Macromolecule #3: NADH dehydrogenase [ubiquinone] iron-sulfur protein 3
+Macromolecule #4: NADH dehydrogenase [ubiquinone] iron-sulfur protein 2
+Macromolecule #5: NADH dehydrogenase [ubiquinone] flavoprotein 2, mitochondrial
+Macromolecule #6: NADH dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial
+Macromolecule #7: NADH dehydrogenase [ubiquinone] iron-sulfur protein 1, mitochondrial
+Macromolecule #8: NADH-ubiquinone oxidoreductase chain 1
+Macromolecule #9: NADH dehydrogenase [ubiquinone] iron-sulfur protein 8-A, mitochondrial
+Macromolecule #10: NADH-ubiquinone oxidoreductase chain 6
+Macromolecule #11: NADH dehydrogenase subunit 4L
+Macromolecule #12: NADH-ubiquinone oxidoreductase chain 5
+Macromolecule #13: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #14: NADH-ubiquinone oxidoreductase chain 2
+Macromolecule #15: AT3G07480.1
+Macromolecule #16: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 9, mit...
+Macromolecule #17: NADH dehydrogenase [ubiquinone] iron-sulfur protein 4, mitochondrial
+Macromolecule #18: NADH dehydrogenase [ubiquinone] iron-sulfur protein 6, mitochondrial
+Macromolecule #19: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 2
+Macromolecule #20: Acyl carrier protein 1, mitochondrial
+Macromolecule #21: Acyl carrier protein 2, mitochondrial
+Macromolecule #22: Probable NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subun...
+Macromolecule #23: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 6
+Macromolecule #24: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 8-B
+Macromolecule #25: Outer envelope pore protein 16-3, chloroplastic/mitochondrial
+Macromolecule #26: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 13-A
+Macromolecule #27: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 1
+Macromolecule #28: At2g46540/F11C10.23
+Macromolecule #29: Transmembrane protein
+Macromolecule #30: Excitatory amino acid transporter
+Macromolecule #31: NADH dehydrogenase [ubiquinone] iron-sulfur protein 5-B
+Macromolecule #32: At4g16450
+Macromolecule #33: ESSS subunit of NADH:ubiquinone oxidoreductase (Complex I) protein
+Macromolecule #34: P1
+Macromolecule #35: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 2
+Macromolecule #36: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 3-A
+Macromolecule #37: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 8, mito...
+Macromolecule #38: B15 -- 1 beta subcomplex subunit 4
+Macromolecule #39: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 9
+Macromolecule #40: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 7
+Macromolecule #41: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 10-B
+Macromolecule #42: Probable NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 12
+Macromolecule #43: Uncharacterized protein At1g67785
+Macromolecule #44: Uncharacterized protein At2g27730, mitochondrial
+Macromolecule #45: Gamma carbonic anhydrase-like 2, mitochondrial
+Macromolecule #46: Gamma carbonic anhydrase 2, mitochondrial
+Macromolecule #47: Gamma carbonic anhydrase 1, mitochondrial
+Macromolecule #48: Probable mitochondrial-processing peptidase subunit alpha-1, mito...
+Macromolecule #49: Probable mitochondrial-processing peptidase subunit beta, mitocho...
+Macromolecule #50: Cytochrome b
+Macromolecule #51: Cytochrome b-c1 complex subunit Rieske-1, mitochondrial
+Macromolecule #52: Cytochrome c1 2, heme protein, mitochondrial
+Macromolecule #53: Cytochrome b-c1 complex subunit 7-2, mitochondrial
+Macromolecule #54: Cytochrome b-c1 complex subunit 8-1, mitochondrial
+Macromolecule #55: Cytochrome b-c1 complex subunit 6-1, mitochondrial
+Macromolecule #56: Cytochrome b-c1 complex subunit 9, mitochondrial
+Macromolecule #57: Cytochrome b-c1 complex subunit 10, mitochondrial
+Macromolecule #58: IRON/SULFUR CLUSTER
+Macromolecule #59: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #60: FLAVIN MONONUCLEOTIDE
+Macromolecule #61: Ubiquinone-9
+Macromolecule #62: FE (III) ION
+Macromolecule #63: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
+Macromolecule #64: ZINC ION
+Macromolecule #65: S-[2-({N-[(2R)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
+Macromolecule #66: CROTONYL COENZYME A
+Macromolecule #67: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #68: 2,3-DIMETHOXY-5-METHYL-6-(3,11,15,19-TETRAMETHYL-EICOSA-2,6,10,14...
+Macromolecule #69: UBIQUINONE-7
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.18 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - #0 - Film type ID: 1 / Support film - #0 - Material: CARBON / Support film - #0 - topology: CONTINUOUS / Support film - #1 - Film type ID: 2 / Support film - #1 - Material: GRAPHENE / Support film - #1 - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 70 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 215000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8bq6: |
 Movie
Movie Controller
Controller






































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)