+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | RecBCD in complex with the phage protein gp5.9 | ||||||||||||
 Map data Map data | Final cryoEM map of the RecBCD-gp5.9 complex | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Homologous recombination / DNA repair / phage / Helicase / Nuclease / Inhibitor / Protein complex / Enzyme / DNA mimic / DNA BINDING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationexodeoxyribonuclease V / exodeoxyribonuclease V activity / exodeoxyribonuclease V complex / clearance of foreign intracellular DNA / DNA 5'-3' helicase / single-stranded DNA helicase activity / recombinational repair / 3'-5' DNA helicase activity / DNA 3'-5' helicase / DNA helicase activity ...exodeoxyribonuclease V / exodeoxyribonuclease V activity / exodeoxyribonuclease V complex / clearance of foreign intracellular DNA / DNA 5'-3' helicase / single-stranded DNA helicase activity / recombinational repair / 3'-5' DNA helicase activity / DNA 3'-5' helicase / DNA helicase activity / helicase activity / response to radiation / double-strand break repair via homologous recombination / DNA recombination / 5'-3' DNA helicase activity / symbiont-mediated suppression of host innate immune response / DNA damage response / magnesium ion binding / ATP hydrolysis activity / DNA binding / ATP binding / cytosol Similarity search - Function | ||||||||||||
| Biological species |    Escherichia phage T7 (virus) Escherichia phage T7 (virus) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | ||||||||||||
 Authors Authors | Wilkinson M / Wilkinson OJ / Feyerherm C / Fletcher EE / Wigley DB / Dillingham MS | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Structures of RecBCD in complex with phage-encoded inhibitor proteins reveal distinctive strategies for evasion of a bacterial immunity hub. Authors: Martin Wilkinson / Oliver J Wilkinson / Connie Feyerherm / Emma E Fletcher / Dale B Wigley / Mark S Dillingham /  Abstract: Following infection of bacterial cells, bacteriophage modulate double-stranded DNA break repair pathways to protect themselves from host immunity systems and prioritise their own recombinases. Here, ...Following infection of bacterial cells, bacteriophage modulate double-stranded DNA break repair pathways to protect themselves from host immunity systems and prioritise their own recombinases. Here, we present biochemical and structural analysis of two phage proteins, gp5.9 and Abc2, which target the DNA break resection complex RecBCD. These exemplify two contrasting mechanisms for control of DNA break repair in which the RecBCD complex is either inhibited or co-opted for the benefit of the invading phage. Gp5.9 completely inhibits RecBCD by preventing it from binding to DNA. The RecBCD-gp5.9 structure shows that gp5.9 acts by substrate mimicry, binding predominantly to the RecB arm domain and competing sterically for the DNA binding site. Gp5.9 adopts a parallel coiled-coil architecture that is unprecedented for a natural DNA mimic protein. In contrast, binding of Abc2 does not substantially affect the biochemical activities of isolated RecBCD. The RecBCD-Abc2 structure shows that Abc2 binds to the Chi-recognition domains of the RecC subunit in a position that might enable it to mediate the loading of phage recombinases onto its single-stranded DNA products. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15803.map.gz emd_15803.map.gz | 39.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15803-v30.xml emd-15803-v30.xml emd-15803.xml emd-15803.xml | 27.7 KB 27.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15803.png emd_15803.png | 106.3 KB | ||
| Filedesc metadata |  emd-15803.cif.gz emd-15803.cif.gz | 8.8 KB | ||
| Others |  emd_15803_additional_1.map.gz emd_15803_additional_1.map.gz emd_15803_half_map_1.map.gz emd_15803_half_map_1.map.gz emd_15803_half_map_2.map.gz emd_15803_half_map_2.map.gz | 39 MB 33 MB 33 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15803 http://ftp.pdbj.org/pub/emdb/structures/EMD-15803 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15803 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15803 | HTTPS FTP |
-Related structure data
| Related structure data |  8b1rMC  8b1tC  8b1uC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15803.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15803.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final cryoEM map of the RecBCD-gp5.9 complex | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
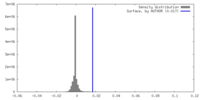
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Additional map from focus refinement around the phage...
| File | emd_15803_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Additional map from focus refinement around the phage protein, used to help model building | ||||||||||||
| Projections & Slices |
| ||||||||||||
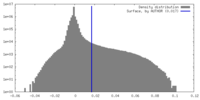
| Density Histograms |
-Half map: halfmap1
| File | emd_15803_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
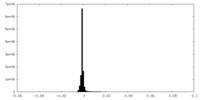
| Density Histograms |
-Half map: halfmap2
| File | emd_15803_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
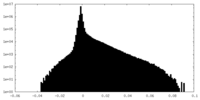
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of gp5.9 bound to RecBCD
| Entire | Name: Complex of gp5.9 bound to RecBCD |
|---|---|
| Components |
|
-Supramolecule #1: Complex of gp5.9 bound to RecBCD
| Supramolecule | Name: Complex of gp5.9 bound to RecBCD / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: t7 gp5.9 purified separately and then mixed with RecBCD complex |
|---|
-Supramolecule #2: RecBCD enzyme subunit RecB, RecC and RecD
| Supramolecule | Name: RecBCD enzyme subunit RecB, RecC and RecD / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Probable RecBCD inhibitor gp5.9
| Supramolecule | Name: Probable RecBCD inhibitor gp5.9 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:   Escherichia phage T7 (virus) Escherichia phage T7 (virus) |
-Macromolecule #1: RecBCD enzyme subunit RecB
| Macromolecule | Name: RecBCD enzyme subunit RecB / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: exodeoxyribonuclease V |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 134.110641 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSDVAETLDP LRLPLQGERL IEASAGTGKT FTIAALYLRL LLGLGGSAAF PRPLTVEELL VVTFTEAATA ELRGRIRSNI HELRIACLR ETTDNPLYER LLEEIDDKAQ AAQWLLLAER QMDEAAVFTI HGFCQRMLNL NAFESGMLFE QQLIEDESLL R YQACADFW ...String: MSDVAETLDP LRLPLQGERL IEASAGTGKT FTIAALYLRL LLGLGGSAAF PRPLTVEELL VVTFTEAATA ELRGRIRSNI HELRIACLR ETTDNPLYER LLEEIDDKAQ AAQWLLLAER QMDEAAVFTI HGFCQRMLNL NAFESGMLFE QQLIEDESLL R YQACADFW RRHCYPLPRE IAQVVFETWK GPQALLRDIN RYLQGEAPVI KAPPPDDETL ASRHAQIVAR IDTVKQQWRD AV GELDALI ESSGIDRRKF NRSNQAKWID KISAWAEEET NSYQLPESLE KFSQRFLEDR TKAGGETPRH PLFEAIDQLL AEP LSIRDL VITRALAEIR ETVAREKRRR GELGFDDMLS RLDSALRSES GEVLAAAIRT RFPVAMIDEF QDTDPQQYRI FRRI WHHQP ETALLLIGDP KQAIYAFRGA DIFTYMKARS EVHAHYTLDT NWRSAPGMVN SVNKLFSQTD DAFMFREIPF IPVKS AGKN QALRFVFKGE TQPAMKMWLM EGESCGVGDY QSTMAQVCAA QIRDWLQAGQ RGEALLMNGD DARPVRASDI SVLVRS RQE AAQVRDALTL LEIPSVYLSN RDSVFETLEA QEMLWLLQAV MTPERENTLR SALATSMMGL NALDIETLNN DEHAWDV VV EEFDGYRQIW RKRGVMPMLR ALMSARNIAE NLLATAGGER RLTDILHISE LLQEAGTQLE SEHALVRWLS QHILEPDS N ASSQQMRLES DKHLVQIVTI HKSKGLEYPL VWLPFITNFR VQEQAFYHDR HSFEAVLDLN AAPESVDLAE AERLAEDLR LLYVALTRSV WHCSLGVAPL VRRRGDKKGD TDVHQSALGR LLQKGEPQDA AGLRTCIEAL CDDDIAWQTA QTGDNQPWQV NDVSTAELN AKTLQRLPGD NWRVTSYSGL QQRGHGIAQD LMPRLDVDAA GVASVVEEPT LTPHQFPRGA SPGTFLHSLF E DLDFTQPV DPNWVREKLE LGGFESQWEP VLTEWITAVL QAPLNETGVS LSQLSARNKQ VEMEFYLPIS EPLIASQLDT LI RQFDPLS AGCPPLEFMQ VRGMLKGFID LVFRHEGRYY LLDYKSNWLG EDSSAYTQQA MAAAMQAHRY DLQYQLYTLA LHR YLRHRI ADYDYEHHFG GVIYLFLRGV DKEHPQQGIY TTRPNAGLIA LMDEMFAGMT LEEA UniProtKB: RecBCD enzyme subunit RecB |
-Macromolecule #2: RecBCD enzyme subunit RecC
| Macromolecule | Name: RecBCD enzyme subunit RecC / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: exodeoxyribonuclease V |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 128.974102 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLRVYHSNRL DVLEALMEFI VERERLDDPF EPEMILVQST GMAQWLQMTL SQKFGIAANI DFPLPASFIW DMFVRVLPEI PKESAFNKQ SMSWKLMTLL PQLLEREDFT LLRHYLTDDS DKRKLFQLSS KAADLFDQYL VYRPDWLAQW ETGHLVEGLG E AQAWQAPL ...String: MLRVYHSNRL DVLEALMEFI VERERLDDPF EPEMILVQST GMAQWLQMTL SQKFGIAANI DFPLPASFIW DMFVRVLPEI PKESAFNKQ SMSWKLMTLL PQLLEREDFT LLRHYLTDDS DKRKLFQLSS KAADLFDQYL VYRPDWLAQW ETGHLVEGLG E AQAWQAPL WKALVEYTHQ LGQPRWHRAN LYQRFIETLE SATTCPPGLP SRVFICGISA LPPVYLQALQ ALGKHIEIHL LF TNPCRYY WGDIKDPAYL AKLLTRQRRH SFEDRELPLF RDSENAGQLF NSDGEQDVGN PLLASWGKLG RDYIYLLSDL ESS QELDAF VDVTPDNLLH NIQSDILELE NRAVAGVNIE EFSRSDNKRP LDPLDSSITF HVCHSPQREV EVLHDRLLAM LEED PTLTP RDIIVMVADI DSYSPFIQAV FGSAPADRYL PYAISDRRAR QSHPVLEAFI SLLSLPDSRF VSEDVLALLD VPVLA ARFD ITEEGLRYLR QWVNESGIRW GIDDDNVREL ELPATGQHTW RFGLTRMLLG YAMESAQGEW QSVLPYDESS GLIAEL VGH LASLLMQLNI WRRGLAQERP LEEWLPVCRD MLNAFFLPDA ETEAAMTLIE QQWQAIIAEG LGAQYGDAVP LSLLRDE LA QRLDQERISQ RFLAGPVNIC TLMPMRSIPF KVVCLLGMND GVYPRQLAPL GFDLMSQKPK RGDRSRRDDD RYLFLEAL I SAQQKLYISY IGRSIQDNSE RFPSVLVQEL IDYIGQSHYL PGDEALNCDE SEARVKAHLT CLHTRMPFDP QNYQPGERQ SYAREWLPAA SQAGKAHSEF VQPLPFTLPE TVPLETLQRF WAHPVRAFFQ MRLQVNFRTE DSEIPDTEPF ILEGLSRYQI NQQLLNALV EQDDAERLFR RFRAAGDLPY GAFGEIFWET QCQEMQQLAD RVIACRQPGQ SMEIDLACNG VQITGWLPQV Q PDGLLRWR PSLLSVAQGM QLWLEHLVYC ASGGNGESRL FLRKDGEWRF PPLAAEQALH YLSQLIEGYR EGMSAPLLVL PE SGGAWLK TCYDAQNDAM LDDDSTLQKA RTKFLQAYEG NMMVRGEGDD IWYQRLWRQL TPETMEAIVE QSQRFLLPLF RFN QS UniProtKB: RecBCD enzyme subunit RecC |
-Macromolecule #3: RecBCD enzyme subunit RecD
| Macromolecule | Name: RecBCD enzyme subunit RecD / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: exodeoxyribonuclease V |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 66.990367 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKLQKQLLEA VEHKQLRPLD VQFALTVAGD EHPAVTLAAA LLSHDAGEGH VCLPLSRLEN NEASHPLLAT CVSEIGELQN WEECLLASQ AVSRGDEPTP MILCGDRLYL NRMWCNERTV ARFFNEVNHA IEVDEALLAQ TLDKLFPVSD EINWQKVAAA V ALTRRISV ...String: MKLQKQLLEA VEHKQLRPLD VQFALTVAGD EHPAVTLAAA LLSHDAGEGH VCLPLSRLEN NEASHPLLAT CVSEIGELQN WEECLLASQ AVSRGDEPTP MILCGDRLYL NRMWCNERTV ARFFNEVNHA IEVDEALLAQ TLDKLFPVSD EINWQKVAAA V ALTRRISV ISGGPGTGKT TTVAKLLAAL IQMADGERCR IRLAAPTGKA AARLTESLGK ALRQLPLTDE QKKRIPEDAS TL HRLLGAQ PGSQRLRHHA GNPLHLDVLV VDEASMIDLP MMSRLIDALP DHARVIFLGD RDQLASVEAG AVLGDICAYA NAG FTAERA RQLSRLTGTH VPAGTGTEAA SLRDSLCLLQ KSYRFGSDSG IGQLAAAINR GDKTAVKTVF QQDFTDIEKR LLQS GEDYI AMLEEALAGY GRYLDLLQAR AEPDLIIQAF NEYQLLCALR EGPFGVAGLN ERIEQFMQQK RKIHRHPHSR WYEGR PVMI ARNDSALGLF NGDIGIALDR GQGTRVWFAM PDGNIKSVQP SRLPEHETTW AMTVHKSQGS EFDHAALILP SQRTPV VTR ELVYTAVTRA RRRLSLYADE RILSAAIATR TERRSGLAAL FSSRE UniProtKB: RecBCD enzyme subunit RecD |
-Macromolecule #4: Probable RecBCD inhibitor gp5.9
| Macromolecule | Name: Probable RecBCD inhibitor gp5.9 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage T7 (virus) Escherichia phage T7 (virus) |
| Molecular weight | Theoretical: 6.049601 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSRDLVTIPR DVWNDIQGYI DSLERENDSL KNQLMEADEY VAELEEKLNG TS UniProtKB: Probable RecBCD inhibitor gp5.9 |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: GRAPHENE OXIDE Details: Mixture of graphene oxide with 0.3 mM DDM detergent applied directly to grids twice before application of sample | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: FEI VITROBOT MARK IV / Details: 1.5s blot time. | |||||||||||||||
| Details | RecBCD mixed with T7 gp5.9 protein prior to making grids |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 5064 / Average exposure time: 4.3 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)