[English] 日本語
 Yorodumi
Yorodumi- EMDB-1570: CryoEM reconstructions of PV1 complexed with deglycosylated CD155 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1570 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM reconstructions of PV1 complexed with deglycosylated CD155 | |||||||||
 Map data Map data | CryoEM reconstructions of PV1 complexed with deglycosylated CD155 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Poliovirus type1 / poliovirus receptor | |||||||||
| Function / homology |  Function and homology information Function and homology informationsusceptibility to T cell mediated cytotoxicity / susceptibility to natural killer cell mediated cytotoxicity / Nectin/Necl trans heterodimerization / positive regulation of natural killer cell mediated cytotoxicity directed against tumor cell target / cell adhesion mediator activity / symbiont-mediated suppression of host translation initiation / positive regulation of natural killer cell mediated cytotoxicity / negative regulation of natural killer cell mediated cytotoxicity / heterophilic cell-cell adhesion / natural killer cell mediated cytotoxicity ...susceptibility to T cell mediated cytotoxicity / susceptibility to natural killer cell mediated cytotoxicity / Nectin/Necl trans heterodimerization / positive regulation of natural killer cell mediated cytotoxicity directed against tumor cell target / cell adhesion mediator activity / symbiont-mediated suppression of host translation initiation / positive regulation of natural killer cell mediated cytotoxicity / negative regulation of natural killer cell mediated cytotoxicity / heterophilic cell-cell adhesion / natural killer cell mediated cytotoxicity / homophilic cell-cell adhesion / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / cell adhesion molecule binding / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / adherens junction / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / virus receptor activity / channel activity / signaling receptor activity / monoatomic ion transmembrane transport / RNA helicase activity / endocytosis involved in viral entry into host cell / receptor ligand activity / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / focal adhesion / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / cell surface / proteolysis / : / RNA binding / zinc ion binding / ATP binding / membrane / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Human poliovirus 1 Human poliovirus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 8.0 Å | |||||||||
 Authors Authors | Zhang P / Mueller S / Morais MC / Bator-Kelly CM / Bowman VD / Hafenstein S / Wimmer E / Rossmann MG | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2008 Journal: Proc Natl Acad Sci U S A / Year: 2008Title: Crystal structure of CD155 and electron microscopic studies of its complexes with polioviruses. Authors: Ping Zhang / Steffen Mueller / Marc C Morais / Carol M Bator / Valorie D Bowman / Susan Hafenstein / Eckard Wimmer / Michael G Rossmann /  Abstract: When poliovirus (PV) recognizes its receptor, CD155, the virus changes from a 160S to a 135S particle before releasing its genome into the cytoplasm. CD155 is a transmembrane protein with 3 Ig-like ...When poliovirus (PV) recognizes its receptor, CD155, the virus changes from a 160S to a 135S particle before releasing its genome into the cytoplasm. CD155 is a transmembrane protein with 3 Ig-like extracellular domains, D1-D3, where D1 is recognized by the virus. The crystal structure of D1D2 has been determined to 3.5-A resolution and fitted into approximately 8.5-A resolution cryoelectron microscopy reconstructions of the virus-receptor complexes for the 3 PV serotypes. These structures show that, compared with human rhinoviruses, the virus-receptor interactions for PVs have a greater dependence on hydrophobic interactions, as might be required for a virus that can inhabit environments of different pH. The pocket factor was shown to remain in the virus during the first recognition stage. The present structures, when combined with earlier mutational investigations, show that in the subsequent entry stage the receptor moves further into the canyon when at a physiological temperature, thereby expelling the pocket factor and separating the viral subunits to form 135S particles. These results provide a detailed analysis of how a nonenveloped virus can enter its host cell. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1570.map.gz emd_1570.map.gz | 9.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1570-v30.xml emd-1570-v30.xml emd-1570.xml emd-1570.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
| Images |  1570.gif 1570.gif | 40.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1570 http://ftp.pdbj.org/pub/emdb/structures/EMD-1570 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1570 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1570 | HTTPS FTP |
-Related structure data
| Related structure data |  3epcMC  1562C  1563C  3epdC  3epfC  3uroC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1570.map.gz / Format: CCP4 / Size: 38.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1570.map.gz / Format: CCP4 / Size: 38.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM reconstructions of PV1 complexed with deglycosylated CD155 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.65 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
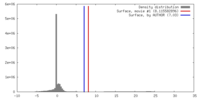
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : CD155-PV1 complex
| Entire | Name: CD155-PV1 complex |
|---|---|
| Components |
|
-Supramolecule #1000: CD155-PV1 complex
| Supramolecule | Name: CD155-PV1 complex / type: sample / ID: 1000 Oligomeric state: 60 copies of CD166 bind to icosahedral protein shell of a PV1 particle Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 11.1 MDa |
-Supramolecule #1: Human poliovirus 1
| Supramolecule | Name: Human poliovirus 1 / type: virus / ID: 1 / Name.synonym: poliovirus type 1 / NCBI-ID: 12080 / Sci species name: Human poliovirus 1 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No / Syn species name: poliovirus type 1 |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Molecular weight | Theoretical: 8.5 MDa |
| Virus shell | Shell ID: 1 / Diameter: 310 Å |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 10mM Tris-HCl, 20mM NaCl |
| Staining | Type: NEGATIVE Details: A small vial of ethane is placed inside a larger liquid nitrogen reservoir. The grid holding a few microliters of the sample is held in place at the bottom of a plunger by the means of fine ...Details: A small vial of ethane is placed inside a larger liquid nitrogen reservoir. The grid holding a few microliters of the sample is held in place at the bottom of a plunger by the means of fine tweezers. Once the ethane in the vial is completely frozen, it needs to be slightly melted. When the liquid ethane is ready, a piece of filter paper is then pressed against the sample to blot of excess buffer, sufficient to leave a thin layer on the grid. After a predetermined time, the filter paper is removed, and the plunger is allowed to drop into the liquid ethane. Once the grid enters the liquid ethane, the sample is rapidly frozen, and the grid is transferred under liquid nitrogen to a storage box immersed liquid nitrogen for later use in the microscope. |
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/T |
|---|---|
| Alignment procedure | Legacy - Astigmatism: live FFT at 200K |
| Image recording | Category: CCD / Film or detector model: KODAK SO-163 FILM / Number real images: 47 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 47190 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.126 µm / Nominal defocus min: 0.857 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 8.0 Å / Resolution method: FSC 0.5 CUT-OFF Details: Final map includes data to 8.0 Ang resolution (fsc 0.5 cut-off), magnification of final map standardized to a map calculated from PV1 atomic coordinates (PDB accession no 2PLV) resulting in ...Details: Final map includes data to 8.0 Ang resolution (fsc 0.5 cut-off), magnification of final map standardized to a map calculated from PV1 atomic coordinates (PDB accession no 2PLV) resulting in final pixel separation of 2.65 Ang |
|---|
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)