+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5515 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Icosahedral reconstruction of filled coxsackievirus A9 capsid | |||||||||
 Map data Map data | icosahedral reconstruction of filled coxsackievirus A9 capsid | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CVA9-integrin / picornavirus / enterovirus / coxsackievirus A9 / integrin | |||||||||
| Biological species |  Human coxsackievirus A9 Human coxsackievirus A9 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 10.34 Å | |||||||||
 Authors Authors | Shakeel S / Seitsonen JJT / Kajander T / Laurinmaki P / Hyypia T / Susi P / Butcher SJ | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2013 Journal: J Virol / Year: 2013Title: Structural and functional analysis of coxsackievirus A9 integrin αvβ6 binding and uncoating. Authors: Shabih Shakeel / Jani J T Seitsonen / Tommi Kajander / Pasi Laurinmäki / Timo Hyypiä / Petri Susi / Sarah J Butcher /  Abstract: Coxsackievirus A9 (CVA9) is an important pathogen of the Picornaviridae family. It utilizes cellular receptors from the integrin αv family for binding to its host cells prior to entry and genome ...Coxsackievirus A9 (CVA9) is an important pathogen of the Picornaviridae family. It utilizes cellular receptors from the integrin αv family for binding to its host cells prior to entry and genome release. Among the integrins tested, it has the highest affinity for αvβ6, which recognizes the arginine-glycine-aspartic acid (RGD) loop present on the C terminus of viral capsid protein, VP1. As the atomic model of CVA9 lacks the RGD loop, we used surface plasmon resonance, electron cryo-microscopy, and image reconstruction to characterize the capsid-integrin interactions and the conformational changes on genome release. We show that the integrin binds to the capsid with nanomolar affinity and that the binding of integrin to the virion does not induce uncoating, thereby implying that further steps are required for release of the genome. Electron cryo-tomography and single-particle image reconstruction revealed variation in the number and conformation of the integrins bound to the capsid, with the integrin footprint mapping close to the predicted site for the exposed RGD loop on VP1. Comparison of empty and RNA-filled capsid reconstructions showed that the capsid undergoes conformational changes when the genome is released, so that the RNA-capsid interactions in the N termini of VP1 and VP4 are lost, VP4 is removed, and the capsid becomes more porous, as has been reported for poliovirus 1, human rhinovirus 2, enterovirus 71, and coxsackievirus A7. These results are important for understanding the structural basis of integrin binding to CVA9 and the molecular events leading to CVA9 cell entry and uncoating. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5515.map.gz emd_5515.map.gz | 20.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5515-v30.xml emd-5515-v30.xml emd-5515.xml emd-5515.xml | 8.2 KB 8.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5515_1.jpg emd_5515_1.jpg | 163 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5515 http://ftp.pdbj.org/pub/emdb/structures/EMD-5515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5515 | HTTPS FTP |
-Related structure data
| Related structure data |  5512C  5514C  5516C  5517C  5519C  3j2jC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5515.map.gz / Format: CCP4 / Size: 120.1 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_5515.map.gz / Format: CCP4 / Size: 120.1 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | icosahedral reconstruction of filled coxsackievirus A9 capsid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.13 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
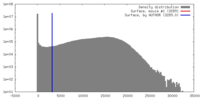
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : filled coxsackievirus A9 capsid
| Entire | Name: filled coxsackievirus A9 capsid |
|---|---|
| Components |
|
-Supramolecule #1000: filled coxsackievirus A9 capsid
| Supramolecule | Name: filled coxsackievirus A9 capsid / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Human coxsackievirus A9
| Supramolecule | Name: Human coxsackievirus A9 / type: virus / ID: 1 / Name.synonym: CVA9 / NCBI-ID: 12067 / Sci species name: Human coxsackievirus A9 / Database: NCBI / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No / Syn species name: CVA9 |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Virus shell | Shell ID: 1 / T number (triangulation number): 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: GATAN CRYOPLUNGE 3 |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Mar 1, 2011 |
| Image recording | Digitization - Scanner: OTHER / Digitization - Sampling interval: 7 µm / Number real images: 56 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 62000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.32 µm / Nominal defocus min: 0.89 µm / Nominal magnification: 62000 |
| Sample stage | Specimen holder: GATAN 626 / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: whole particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 10.34 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: AUTO3DEM / Number images used: 488 |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)