[English] 日本語
 Yorodumi
Yorodumi- EMDB-15451: Sub-tomogram average of TAILS in bovine sperm tail tip singlet mi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
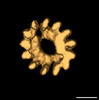
| Title | Sub-tomogram average of TAILS in bovine sperm tail tip singlet microtubules. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Microtubules Flagella Bovine Sperm / UNKNOWN FUNCTION | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 20.0 Å | |||||||||
 Authors Authors | Zabeo D / Hoog JL | |||||||||
| Funding support |  Sweden, 1 items Sweden, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: SPACA9 is a lumenal protein of human ciliary singlet and doublet microtubules. Authors: Miao Gui / Jacob T Croft / Davide Zabeo / Vajradhar Acharya / Justin M Kollman / Thomas Burgoyne / Johanna L Höög / Alan Brown /    Abstract: The cilium-centrosome complex contains triplet, doublet, and singlet microtubules. The lumenal surfaces of each microtubule within this diverse array are decorated by microtubule inner proteins ...The cilium-centrosome complex contains triplet, doublet, and singlet microtubules. The lumenal surfaces of each microtubule within this diverse array are decorated by microtubule inner proteins (MIPs). Here, we used single-particle cryo-electron microscopy methods to build atomic models of two types of human ciliary microtubule: the doublet microtubules of multiciliated respiratory cells and the distal singlet microtubules of monoflagellated human spermatozoa. We discover that SPACA9 is a polyspecific MIP capable of binding both microtubule types. SPACA9 forms intralumenal striations in the B tubule of respiratory doublet microtubules and noncontinuous spirals in sperm singlet microtubules. By acquiring new and reanalyzing previous cryo-electron tomography data, we show that SPACA9-like intralumenal striations are common features of different microtubule types in animal cilia. Our structures provide detailed references to help rationalize ciliopathy-causing mutations and position cryo-EM as a tool for the analysis of samples obtained directly from ciliopathy patients. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15451.map.gz emd_15451.map.gz | 2.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15451-v30.xml emd-15451-v30.xml emd-15451.xml emd-15451.xml | 12.9 KB 12.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15451.png emd_15451.png | 55.7 KB | ||
| Filedesc metadata |  emd-15451.cif.gz emd-15451.cif.gz | 3.8 KB | ||
| Others |  emd_15451_half_map_1.map.gz emd_15451_half_map_1.map.gz emd_15451_half_map_2.map.gz emd_15451_half_map_2.map.gz | 2.9 MB 2.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15451 http://ftp.pdbj.org/pub/emdb/structures/EMD-15451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15451 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15451.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15451.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.37 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_15451_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_15451_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SPACA9 inner microtubule decoration.
| Entire | Name: SPACA9 inner microtubule decoration. |
|---|---|
| Components |
|
-Supramolecule #1: SPACA9 inner microtubule decoration.
| Supramolecule | Name: SPACA9 inner microtubule decoration. / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.64 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 6.0 µm / Nominal defocus min: 4.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: OTHER / Software - Name: PEET / Number subtomograms used: 14549 |
|---|---|
| Extraction | Number tomograms: 10 / Number images used: 14549 / Software - Name: PEET |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: PEET |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































