+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13999 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tomogram of skeletal sarcomere I-band after FIB-milling | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Skeletal muscle / Sarcomere / I-band / CONTRACTILE PROTEIN | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | electron tomography / cryo EM | ||||||||||||||||||
 Authors Authors | Wang Z / Raunser S | ||||||||||||||||||
| Funding support | European Union, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structures from intact myofibrils reveal mechanism of thin filament regulation through nebulin. Authors: Zhexin Wang / Michael Grange / Sabrina Pospich / Thorsten Wagner / Ay Lin Kho / Mathias Gautel / Stefan Raunser /   Abstract: In skeletal muscle, nebulin stabilizes and regulates the length of thin filaments, but the underlying mechanism remains nebulous. In this work, we used cryo-electron tomography and subtomogram ...In skeletal muscle, nebulin stabilizes and regulates the length of thin filaments, but the underlying mechanism remains nebulous. In this work, we used cryo-electron tomography and subtomogram averaging to reveal structures of native nebulin bound to thin filaments within intact sarcomeres. This in situ reconstruction provided high-resolution details of the interaction between nebulin and actin, demonstrating the stabilizing role of nebulin. Myosin bound to the thin filaments exhibited different conformations of the neck domain, highlighting its inherent structural variability in muscle. Unexpectedly, nebulin did not interact with myosin or tropomyosin, but it did interact with a troponin T linker through two potential binding motifs on nebulin, explaining its regulatory role. Our structures support the role of nebulin as a thin filament "molecular ruler" and provide a molecular basis for studying nemaline myopathies. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13999.map.gz emd_13999.map.gz | 1.2 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13999-v30.xml emd-13999-v30.xml emd-13999.xml emd-13999.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13999.png emd_13999.png | 149.4 KB | ||
| Filedesc metadata |  emd-13999.cif.gz emd-13999.cif.gz | 4.1 KB | ||
| Others |  emd_13999_additional_1.map.gz emd_13999_additional_1.map.gz | 1.2 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13999 http://ftp.pdbj.org/pub/emdb/structures/EMD-13999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13999 | HTTPS FTP |
-Validation report
| Summary document |  emd_13999_validation.pdf.gz emd_13999_validation.pdf.gz | 486.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_13999_full_validation.pdf.gz emd_13999_full_validation.pdf.gz | 485.8 KB | Display | |
| Data in XML |  emd_13999_validation.xml.gz emd_13999_validation.xml.gz | 5.1 KB | Display | |
| Data in CIF |  emd_13999_validation.cif.gz emd_13999_validation.cif.gz | 5.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13999 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13999 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13999 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13999 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13999.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13999.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.716 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_13999_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
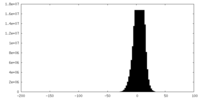
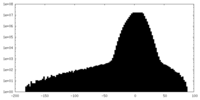
| Density Histograms |
- Sample components
Sample components
-Entire : Mouse psoas muscle myofibrils
| Entire | Name: Mouse psoas muscle myofibrils |
|---|---|
| Components |
|
-Supramolecule #1: Mouse psoas muscle myofibrils
| Supramolecule | Name: Mouse psoas muscle myofibrils / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | tissue |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 / Focused ion beam - Current: 0.05 / Focused ion beam - Duration: 60 / Focused ion beam - Temperature: 90 K / Focused ion beam - Initial thickness: 2000 / Focused ion beam - Final thickness: 100 Focused ion beam - Details: The value given for _em_focused_ion_beam.instrument is FEI Aquilos. This is not in a list of allowed values {'DB235', 'OTHER'} so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: COUNTING / Average electron dose: 3.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 5.5 µm / Calibrated defocus min: 2.5 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.5 µm / Nominal defocus min: 2.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Number images used: 37 |
|---|
-Atomic model buiding 1
| Refinement | Overall B value: 250 |
|---|
 Movie
Movie Controller
Controller
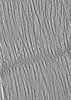

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)