[English] 日本語
 Yorodumi
Yorodumi- EMDB-13748: Preliminary Single particle cryo-EM reconstruction of a 40-mer as... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13748 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Preliminary Single particle cryo-EM reconstruction of a 40-mer assembly of Hsp26 purified from yeast. | |||||||||
 Map data Map data | Preliminary WT map of Hsp26 purified from yeast | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasmic stress granule / : / cellular response to heat / protein folding / mRNA binding / mitochondrion / identical protein binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Peters C / Weinkauf S / Buchner J | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Phosphorylation activates the yeast small heat shock protein Hsp26 by weakening domain contacts in the oligomer ensemble. Authors: Moritz Mühlhofer / Carsten Peters / Thomas Kriehuber / Marina Kreuzeder / Pamina Kazman / Natalia Rodina / Bernd Reif / Martin Haslbeck / Sevil Weinkauf / Johannes Buchner /  Abstract: Hsp26 is a small heat shock protein (sHsp) from S. cerevisiae. Its chaperone activity is activated by oligomer dissociation at heat shock temperatures. Hsp26 contains 9 phosphorylation sites in ...Hsp26 is a small heat shock protein (sHsp) from S. cerevisiae. Its chaperone activity is activated by oligomer dissociation at heat shock temperatures. Hsp26 contains 9 phosphorylation sites in different structural elements. Our analysis of phospho-mimetic mutations shows that phosphorylation activates Hsp26 at permissive temperatures. The cryo-EM structure of the Hsp26 40mer revealed contacts between the conserved core domain of Hsp26 and the so-called thermosensor domain in the N-terminal part of the protein, which are targeted by phosphorylation. Furthermore, several phosphorylation sites in the C-terminal extension, which link subunits within the oligomer, are sensitive to the introduction of negative charges. In all cases, the intrinsic inhibition of chaperone activity is relieved and the N-terminal domain becomes accessible for substrate protein binding. The weakening of domain interactions within and between subunits by phosphorylation to activate the chaperone activity in response to proteotoxic stresses independent of heat stress could be a general regulation principle of sHsps. #1:  Journal: Structure / Year: 2006 Journal: Structure / Year: 2006Title: Multiple distinct assemblies reveal conformational flexibility in the small heat shock protein Hsp26. Authors: White HE / Orlova EV / Chen S / Wang L / Ignatiou A / Gowen B / Stromer T / Franzmann TM / Haslbeck M / Buchner J / Saibil HR | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13748.map.gz emd_13748.map.gz | 39.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13748-v30.xml emd-13748-v30.xml emd-13748.xml emd-13748.xml | 13.9 KB 13.9 KB | Display Display |  EMDB header EMDB header |
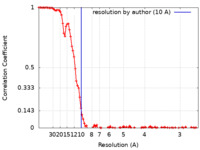
| FSC (resolution estimation) |  emd_13748_fsc.xml emd_13748_fsc.xml | 13 KB | Display |  FSC data file FSC data file |
| Images |  emd_13748.png emd_13748.png | 71.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13748 http://ftp.pdbj.org/pub/emdb/structures/EMD-13748 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13748 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13748 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13748.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13748.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Preliminary WT map of Hsp26 purified from yeast | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.33 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 40mer complex of WT Hsp26
| Entire | Name: 40mer complex of WT Hsp26 |
|---|---|
| Components |
|
-Supramolecule #1: 40mer complex of WT Hsp26
| Supramolecule | Name: 40mer complex of WT Hsp26 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 955.2 KDa |
-Macromolecule #1: Hsp26
| Macromolecule | Name: Hsp26 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MSFNSPFFDF FDNINNEVDA FNRLLGEGGL RGYAPRRQLA NTPAKDSTGK EVARPNNYAG ALYDPRDET LDDWFDNDLS LFPSGFGFPR SVAVPVDILD HDNNYELKVV VPGVKSKKDI D IEYHQNKN QILVSGEIPS TLNEESKDKV KVKESSSGKF KRVITLPDYP ...String: MSFNSPFFDF FDNINNEVDA FNRLLGEGGL RGYAPRRQLA NTPAKDSTGK EVARPNNYAG ALYDPRDET LDDWFDNDLS LFPSGFGFPR SVAVPVDILD HDNNYELKVV VPGVKSKKDI D IEYHQNKN QILVSGEIPS TLNEESKDKV KVKESSSGKF KRVITLPDYP GVDADNIKAD YA NGVLTLT VPKLKPQKDG KNHVKKIEVS SQESWGN |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 294 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 1170 / Average exposure time: 8.0 sec. / Average electron dose: 140.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)