[English] 日本語
 Yorodumi
Yorodumi- EMDB-12947: Cryo-EM structure of tridecameric SlyB from Escherichia coli K12 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of tridecameric SlyB from Escherichia coli K12 | |||||||||||||||
 Map data Map data | SlyB13 | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Outer membrane chaperon / 2TM glycine zipper / outer membrane lipoprotein SlyB / LPS-LP binding protein / MEMBRANE PROTEIN | |||||||||||||||
| Biological species |   | |||||||||||||||
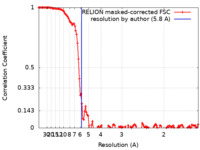
| Method | single particle reconstruction / cryo EM / Resolution: 5.8 Å | |||||||||||||||
 Authors Authors | Nguyen VS / Remaut H | |||||||||||||||
| Funding support |  Belgium, 4 items Belgium, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: SlyB encapsulates outer membrane proteins in stress-induced lipid nanodomains Authors: Janssens A / Nguyen VS / Cecil AJ / Van der Verren SE / Timmerman E / Deghelt M / Pak AJ / Collet JF / Impens F / Remaut H | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12947.map.gz emd_12947.map.gz | 18.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12947-v30.xml emd-12947-v30.xml emd-12947.xml emd-12947.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12947_fsc.xml emd_12947_fsc.xml | 12.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_12947.png emd_12947.png | 129.3 KB | ||
| Filedesc metadata |  emd-12947.cif.gz emd-12947.cif.gz | 4.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12947 http://ftp.pdbj.org/pub/emdb/structures/EMD-12947 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12947 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12947 | HTTPS FTP |
-Validation report
| Summary document |  emd_12947_validation.pdf.gz emd_12947_validation.pdf.gz | 377.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12947_full_validation.pdf.gz emd_12947_full_validation.pdf.gz | 377 KB | Display | |
| Data in XML |  emd_12947_validation.xml.gz emd_12947_validation.xml.gz | 13.1 KB | Display | |
| Data in CIF |  emd_12947_validation.cif.gz emd_12947_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12947 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12947 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12947 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12947 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12947.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12947.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SlyB13 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.784 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Outer membrane lipoprotein SlyB
| Entire | Name: Outer membrane lipoprotein SlyB |
|---|---|
| Components |
|
-Supramolecule #1: Outer membrane lipoprotein SlyB
| Supramolecule | Name: Outer membrane lipoprotein SlyB / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Tricameric SlyB was purified from overexpression of SlyB-TEV-HIS in E. coli using using 2-step Ni-IMAC purification. |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 185 KDa |
-Macromolecule #1: Outer membrane lipoprotein SlyB
| Macromolecule | Name: Outer membrane lipoprotein SlyB / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: (PLM)CVNNDTLSG DVYTASEAKQ VQNVSYGTIV NVRPVQIQGG DDSNVIGAIG GAVLGGFLGN TVGGGTGRSL ATAAGAVAGG VAGQGVQSAM NKTQGVELEI RKDDGNTIMV VQKQGNTRFS PGQRVVLASN GSQVTVSPRG GGENLYFQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.04 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Support film - Material: GRAPHENE OXIDE / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: GATAN CRYOPLUNGE 3 / Details: Back-blotting for 4 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 6352 / Average exposure time: 3.0 sec. / Average electron dose: 61.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.55 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)