[English] 日本語
 Yorodumi
Yorodumi- EMDB-11443: Human pre-40S particle purified using RIO1(kd)-StHA as bait - Str... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11443 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human pre-40S particle purified using RIO1(kd)-StHA as bait - Structural State B, Head | |||||||||
 Map data Map data | Human pre-40S particle purified using RIO1(kd)-StHA as bait - Structural State B, Head | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
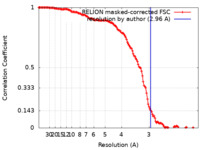
| Method | single particle reconstruction / cryo EM / Resolution: 2.96 Å | |||||||||
 Authors Authors | Plassart L / Shayan R / Plisson-Chastang C | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2021 Journal: Elife / Year: 2021Title: The final step of 40S ribosomal subunit maturation is controlled by a dual key lock. Authors: Laura Plassart / Ramtin Shayan / Christian Montellese / Dana Rinaldi / Natacha Larburu / Carole Pichereaux / Carine Froment / Simon Lebaron / Marie-Françoise O'Donohue / Ulrike Kutay / ...Authors: Laura Plassart / Ramtin Shayan / Christian Montellese / Dana Rinaldi / Natacha Larburu / Carole Pichereaux / Carine Froment / Simon Lebaron / Marie-Françoise O'Donohue / Ulrike Kutay / Julien Marcoux / Pierre-Emmanuel Gleizes / Celia Plisson-Chastang /   Abstract: Preventing premature interaction of pre-ribosomes with the translation apparatus is essential for translational accuracy. Hence, the final maturation step releasing functional 40S ribosomal subunits, ...Preventing premature interaction of pre-ribosomes with the translation apparatus is essential for translational accuracy. Hence, the final maturation step releasing functional 40S ribosomal subunits, namely processing of the 18S ribosomal RNA 3' end, is safeguarded by the protein DIM2, which both interacts with the endoribonuclease NOB1 and masks the rRNA cleavage site. To elucidate the control mechanism that unlocks NOB1 activity, we performed cryo-electron microscopy analysis of late human pre-40S particles purified using a catalytically inactive form of the ATPase RIO1. These structures, together with in vivo and in vitro functional analyses, support a model in which ATP-loaded RIO1 cooperates with ribosomal protein RPS26/eS26 to displace DIM2 from the 18S rRNA 3' end, thereby triggering final cleavage by NOB1; release of ADP then leads to RIO1 dissociation from the 40S subunit. This dual key lock mechanism requiring RIO1 and RPS26 guarantees the precise timing of pre-40S particle conversion into translation-competent ribosomal subunits. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11443.map.gz emd_11443.map.gz | 9.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11443-v30.xml emd-11443-v30.xml emd-11443.xml emd-11443.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11443_fsc.xml emd_11443_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_11443.png emd_11443.png | 178 KB | ||
| Others |  emd_11443_half_map_1.map.gz emd_11443_half_map_1.map.gz emd_11443_half_map_2.map.gz emd_11443_half_map_2.map.gz | 140.9 MB 141 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11443 http://ftp.pdbj.org/pub/emdb/structures/EMD-11443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11443 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11443.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11443.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human pre-40S particle purified using RIO1(kd)-StHA as bait - Structural State B, Head | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #2
| File | emd_11443_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
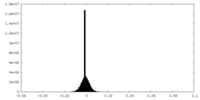
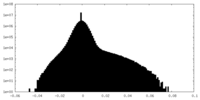
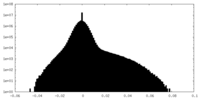
| Density Histograms |
-Half map: #1
| File | emd_11443_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human cytoplasmic late precursor to the small ribosomal subunit, ...
| Entire | Name: human cytoplasmic late precursor to the small ribosomal subunit, purified using RIO1(kd)-StHA as bait. Structural state A (pre 18S rRNA maturation) |
|---|---|
| Components |
|
-Supramolecule #1: human cytoplasmic late precursor to the small ribosomal subunit, ...
| Supramolecule | Name: human cytoplasmic late precursor to the small ribosomal subunit, purified using RIO1(kd)-StHA as bait. Structural state A (pre 18S rRNA maturation) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#35 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Strain: HEK 293 Homo sapiens (human) / Strain: HEK 293 |
| Molecular weight | Theoretical: 1.4 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 291 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 9494 / Average electron dose: 29.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID:  6ek0 |
|---|---|
| Refinement | Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)