+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10730 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Conformation2_Mcm2,5-ssDNA_MultiBody | |||||||||
 Map data Map data | Conformation2-multibody-Mcm2,5-ssDNA | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
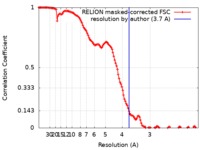
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Baretic D / Jenkyn-Bedford M | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2020 Journal: Mol Cell / Year: 2020Title: Cryo-EM Structure of the Fork Protection Complex Bound to CMG at a Replication Fork. Authors: Domagoj Baretić / Michael Jenkyn-Bedford / Valentina Aria / Giuseppe Cannone / Mark Skehel / Joseph T P Yeeles /  Abstract: The eukaryotic replisome, organized around the Cdc45-MCM-GINS (CMG) helicase, orchestrates chromosome replication. Multiple factors associate directly with CMG, including Ctf4 and the heterotrimeric ...The eukaryotic replisome, organized around the Cdc45-MCM-GINS (CMG) helicase, orchestrates chromosome replication. Multiple factors associate directly with CMG, including Ctf4 and the heterotrimeric fork protection complex (Csm3/Tof1 and Mrc1), which has important roles including aiding normal replication rates and stabilizing stalled forks. How these proteins interface with CMG to execute these functions is poorly understood. Here we present 3 to 3.5 Å resolution electron cryomicroscopy (cryo-EM) structures comprising CMG, Ctf4, and the fork protection complex at a replication fork. The structures provide high-resolution views of CMG-DNA interactions, revealing a mechanism for strand separation, and show Csm3/Tof1 "grip" duplex DNA ahead of CMG via a network of interactions important for efficient replication fork pausing. Although Mrc1 was not resolved in our structures, we determine its topology in the replisome by cross-linking mass spectrometry. Collectively, our work reveals how four highly conserved replisome components collaborate with CMG to facilitate replisome progression and maintain genome stability. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10730.map.gz emd_10730.map.gz | 163.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10730-v30.xml emd-10730-v30.xml emd-10730.xml emd-10730.xml | 17.9 KB 17.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10730_fsc.xml emd_10730_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_10730.png emd_10730.png | 92.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10730 http://ftp.pdbj.org/pub/emdb/structures/EMD-10730 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10730 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10730 | HTTPS FTP |
-Validation report
| Summary document |  emd_10730_validation.pdf.gz emd_10730_validation.pdf.gz | 288.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10730_full_validation.pdf.gz emd_10730_full_validation.pdf.gz | 287.9 KB | Display | |
| Data in XML |  emd_10730_validation.xml.gz emd_10730_validation.xml.gz | 12.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10730 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10730 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10730 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10730 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10730.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10730.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Conformation2-multibody-Mcm2,5-ssDNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.049 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
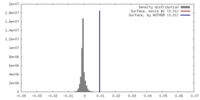
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Conformation 2 of CMG-Csm3-Tof1-Mrc1-Ctf4 with a fork DNA
| Entire | Name: Conformation 2 of CMG-Csm3-Tof1-Mrc1-Ctf4 with a fork DNA |
|---|---|
| Components |
|
-Supramolecule #1: Conformation 2 of CMG-Csm3-Tof1-Mrc1-Ctf4 with a fork DNA
| Supramolecule | Name: Conformation 2 of CMG-Csm3-Tof1-Mrc1-Ctf4 with a fork DNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 / Details: Mcm2,5 domains with bound fragment of ssDNA |
|---|---|
| Molecular weight | Theoretical: 1.4 MDa |
-Supramolecule #2: CMG-Csm3-Tof1-Mrc1-Ctf4
| Supramolecule | Name: CMG-Csm3-Tof1-Mrc1-Ctf4 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1, #3-#7 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Supramolecule #3: DNA
| Supramolecule | Name: DNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism: synthetic construct (others) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 5.0 nm / Pretreatment - Type: PLASMA CLEANING / Details: plasma current of 15 mA | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER Details: Three microlitres of sample was applied on a grid and incubated for 15-30 s at 4 degC before manually blotting with filter paper for 10 s and plunge-freezing in liquid ethane.. | |||||||||||||||||||||
| Details | In vitro reconstitution from individual components: CMG, Csm3-Tof1, Mrc1, Ctf4 and a DNA fork |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-20 / Number grids imaged: 2 / Number real images: 6682 / Average exposure time: 7.0 sec. / Average electron dose: 37.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 3.0 µm / Calibrated defocus min: 0.5 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.4 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)