登録情報 データベース : EMDB / ID : EMD-0781タイトル human PA200-20S complex 細胞 : complex of human PA200-20Sリガンド : x 2種 / 機能・相同性 分子機能 ドメイン・相同性 構成要素
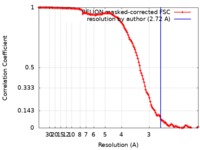
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト)手法 / / 解像度 : 2.72 Å Ouyang S / Hongxin G 資金援助 Organization Grant number 国 National Science Foundation (NSF, China) 31770948
ジャーナル : PLoS Biol / 年 : 2020タイトル : Cryo-EM structures of the human PA200 and PA200-20S complex reveal regulation of proteasome gate opening and two PA200 apertures.著者 : Hongxin Guan / Youwang Wang / Ting Yu / Yini Huang / Mianhuan Li / Abdullah F U H Saeed / Vanja Perčulija / Daliang Li / Jia Xiao / Dongmei Wang / Ping Zhu / Songying Ouyang / 要旨 : Proteasomes are highly abundant and conserved protease complexes that eliminate unwanted proteins in the cells. As a single-chain ATP-independent nuclear proteasome activator, proteasome activator ... Proteasomes are highly abundant and conserved protease complexes that eliminate unwanted proteins in the cells. As a single-chain ATP-independent nuclear proteasome activator, proteasome activator 200 (PA200) associates with 20S core particle to form proteasome complex that catalyzes polyubiquitin-independent degradation of acetylated histones, thus playing a pivotal role in DNA repair and spermatogenesis. Here, we present cryo-electron microscopy (cryo-EM) structures of the human PA200-20S complex and PA200 at 2.72 Å and 3.75 Å, respectively. PA200 exhibits a dome-like architecture that caps 20S and uses its C-terminal YYA (Tyr-Tyr-Ala) to induce the α-ring rearrangements and partial opening of the 20S gate. Our structural data also indicate that PA200 has two openings formed by numerous positively charged residues that respectively bind (5,6)-bisdiphosphoinositol tetrakisphosphate (5,6[PP]2-InsP4) and inositol hexakisphosphate (InsP6) and are likely to be the gates that lead unfolded proteins through PA200 and into the 20S. Besides, our structural analysis of PA200 found that the bromodomain (BRD)-like (BRDL) domain of PA200 shows considerable sequence variation in comparison to other human BRDs, as it contains only 82 residues because of a short ZA loop, and cannot be classified into any of the eight typical human BRD families. Taken together, the results obtained from this study provide important insights into human PA200-induced 20S gate opening for substrate degradation and the opportunities to explore the mechanism for its recognition of H4 histone in acetylation-mediated proteasomal degradation. 履歴 登録 2019年9月9日 - ヘッダ(付随情報) 公開 2020年4月1日 - マップ公開 2020年4月1日 - 更新 2024年3月27日 - 現状 2024年3月27日 処理サイト : PDBj / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト)
Homo sapiens (ヒト) データ登録者
データ登録者 中国, 1件
中国, 1件  引用
引用 ジャーナル: PLoS Biol / 年: 2020
ジャーナル: PLoS Biol / 年: 2020
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_0781.map.gz
emd_0781.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-0781-v30.xml
emd-0781-v30.xml emd-0781.xml
emd-0781.xml EMDBヘッダ
EMDBヘッダ emd_0781_fsc.xml
emd_0781_fsc.xml FSCデータファイル
FSCデータファイル emd_0781.png
emd_0781.png emd-0781.cif.gz
emd-0781.cif.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-0781
http://ftp.pdbj.org/pub/emdb/structures/EMD-0781 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0781
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0781 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_0781.map.gz / 形式: CCP4 / 大きさ: 178 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_0781.map.gz / 形式: CCP4 / 大きさ: 178 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)