+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Wide Pick Filament from Pick's disease brain | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
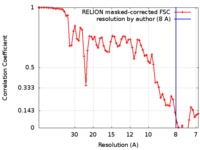
| Method | helical reconstruction / cryo EM / Resolution: 8.0 Å | |||||||||
 Authors Authors | Falcon B / Zhang W / Murzin AG / Murshudov G / Garringer HJ / Vidal R / Crowther RA / Ghetti B / Scheres SHW / Goedert M | |||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Structures of filaments from Pick's disease reveal a novel tau protein fold. Authors: Benjamin Falcon / Wenjuan Zhang / Alexey G Murzin / Garib Murshudov / Holly J Garringer / Ruben Vidal / R Anthony Crowther / Bernardino Ghetti / Sjors H W Scheres / Michel Goedert /   Abstract: The ordered assembly of tau protein into abnormal filamentous inclusions underlies many human neurodegenerative diseases. Tau assemblies seem to spread through specific neural networks in each ...The ordered assembly of tau protein into abnormal filamentous inclusions underlies many human neurodegenerative diseases. Tau assemblies seem to spread through specific neural networks in each disease, with short filaments having the greatest seeding activity. The abundance of tau inclusions strongly correlates with disease symptoms. Six tau isoforms are expressed in the normal adult human brain-three isoforms with four microtubule-binding repeats each (4R tau) and three isoforms that lack the second repeat (3R tau). In various diseases, tau filaments can be composed of either 3R or 4R tau, or of both. Tau filaments have distinct cellular and neuroanatomical distributions, with morphological and biochemical differences suggesting that they may be able to adopt disease-specific molecular conformations. Such conformers may give rise to different neuropathological phenotypes, reminiscent of prion strains. However, the underlying structures are not known. Using electron cryo-microscopy, we recently reported the structures of tau filaments from patients with Alzheimer's disease, which contain both 3R and 4R tau. Here we determine the structures of tau filaments from patients with Pick's disease, a neurodegenerative disorder characterized by frontotemporal dementia. The filaments consist of residues Lys254-Phe378 of 3R tau, which are folded differently from the tau filaments in Alzheimer's disease, establishing the existence of conformers of assembled tau. The observed tau fold in the filaments of patients with Pick's disease explains the selective incorporation of 3R tau in Pick bodies, and the differences in phosphorylation relative to the tau filaments of Alzheimer's disease. Our findings show how tau can adopt distinct folds in the human brain in different diseases, an essential step for understanding the formation and propagation of molecular conformers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0078.map.gz emd_0078.map.gz | 19.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0078-v30.xml emd-0078-v30.xml emd-0078.xml emd-0078.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0078_fsc.xml emd_0078_fsc.xml | 6.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_0078.png emd_0078.png | 38.9 KB | ||
| Masks |  emd_0078_msk_1.map emd_0078_msk_1.map | 22.2 MB |  Mask map Mask map | |
| Others |  emd_0078_half_map_1.map.gz emd_0078_half_map_1.map.gz emd_0078_half_map_2.map.gz emd_0078_half_map_2.map.gz | 16.1 MB 16.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0078 http://ftp.pdbj.org/pub/emdb/structures/EMD-0078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0078 | HTTPS FTP |
-Validation report
| Summary document |  emd_0078_validation.pdf.gz emd_0078_validation.pdf.gz | 384.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0078_full_validation.pdf.gz emd_0078_full_validation.pdf.gz | 383.7 KB | Display | |
| Data in XML |  emd_0078_validation.xml.gz emd_0078_validation.xml.gz | 12.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0078 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0078 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0078 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0078 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0078.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0078.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.45 Å | ||||||||||||||||||||||||||||||||||||
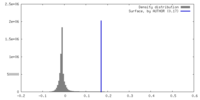
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_0078_msk_1.map emd_0078_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
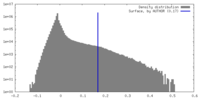
| Density Histograms |
-Half map: #1
| File | emd_0078_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
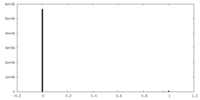
| Density Histograms |
-Half map: #2
| File | emd_0078_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
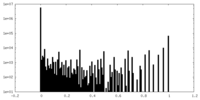
| Density Histograms |
- Sample components
Sample components
-Entire : microtubule-associated protein tau
| Entire | Name: microtubule-associated protein tau |
|---|---|
| Components |
|
-Supramolecule #1: microtubule-associated protein tau
| Supramolecule | Name: microtubule-associated protein tau / type: tissue / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Brain / Tissue: Frontotemporal cortex Homo sapiens (human) / Organ: Brain / Tissue: Frontotemporal cortex |
-Macromolecule #1: Wide Pick Filament from Pick's disease brain
| Macromolecule | Name: Wide Pick Filament from Pick's disease brain / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Sequence | String: KNVKSKIGST ENLKHQPGGG KVQIVYKPVD LSKVTSKCGS LGNIHHKPGG GQVEVKSEKL DFKDRVQSKI GSLDNITHVP GGGNKKIETH KLTF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | tissue |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | Dispersed filaments |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.06 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.8 µm / Nominal defocus min: 1.7 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)